MOLECULAR PHYLOGENY OF THE NERITIDAE (GASTROPODA: NERITIMORPHA) BASED ON THE MITOCHONDRIAL GENES CYTOCHROME OXIDASE I (COI) AND 16S rRNA
Filogenia molecular de la familia Neritidae (Gastropoda: Neritimorpha) con base en los genes mitocondriales citocromo oxidasa I (COI) y 16S rRNA
Palabras clave:
Colombian Caribbean, mDNA, Nerita, Neritina, radiation, mitochondrial genes (en)Nerita, Neritina, Caribe colombiano, genoma mitocondrial, ADNmt, radiación. (es)
Descargas
The family Neritidae has representatives in tropical and subtropical regions that occur in a variety of environments, and its known fossil record dates back to the late Cretaceous. However there have been few studies of molecular phylogeny in this family. We performed a phylogenetic reconstruction of the family Neritidae using the COI (722 bp) and the 16S rRNA (559 bp) regions of the mitochondrial genome. Neighbor-joining, maximum parsimony and Bayesian inference were performed. The best phylogenetic reconstruction was obtained using the COI region, and we consider it an appropriate marker for phylogenetic studies within the group. Consensus analysis (COI +16S rRNA) generally obtained the same tree topologies and confirmed that the genus Nerita is monophyletic. The consensus analysis using parsimony recovered a monophyletic group consisting of the genera Neritina, Septaria, Theodoxus, Puperita, and Clithon, while in the Bayesian analyses Theodoxus is separated from the other genera. The phylogenetic status of the species from the genus Nerita from the Colombian Caribbean generated in this study was consistent with that reported for the genus in previous studies. In the resulting consensus tree obtained using maximum parsimony, we included information on habitat type for each species, to map the evolution by habitat. Species of the family Neritidae possibly have their origin in marine environments, which is consistent with conclusions from previous reports based on anatomical studies.
La familia Neritidae cuenta con representantes en regiones tropicales y subtropicales adaptadas a diferentes ambientes, con un registro fósil que data para finales del Cretáceo. Sin embargo no se han realizado estudios de filogenia molecular en la familia. En este estudio se realizó una reconstrucción filogenética de la familia Neritidae utilizando las regiones COI (722 pb) y 16S rRNA (559 pb) del genoma mitocondrial. Se realizaron análisis de distancias de Neighbor-Joining, Máxima Parsimonia e Inferencia Bayesiana. La mejor reconstrucción filogenética fue mediante la región COI, considerándola un marcador apropiado para realizar estudios filogenéticos dentro del grupo. El consenso de las relaciones filogenéticas (COI+16S rRNA) permitió confirmar que el género Nerita es monofilético. El consenso del análisis de parsimonia reveló un grupo monofilético formado por los géneros Neritina, Septaria, Theodoxus, Puperita y Clithon, mientras que en el análisis bayesiano Theodoxus se encuentra separado de los otros géneros. El resultado en las especies del género Nerita del Caribe colombiano fue consistente con lo reportado para el género en estudios previos. En el árbol resultante del análisis de parsimonia se sobrepuso la información del hábitat de cada especie, para mapear la evolución por hábitat. Se obtuvo como resultado que las especies de la familia Neritidae posiblemente tengan su origen en un ambiente marino, siendo congruente con lo reportado en estudios anatómicos realizados anteriormente.
MOLECULAR PHYLOGENY OF THE NERITIDAE (GASTROPODA: NERITIMORPHA) BASED ON THE MITOCHONDRIAL GENES CYTOCHROME OXIDASE I (COI) AND 16S RRNA
Filogenia molecular de la familia Neritidae (Gastropoda: Neritimorpha) con base en los genes mitocondriales citocromo oxidasa I (COI) y 16S rRNA
JULIAN QUINTERO-GALVIS1, Biólogo; LYDA RAQUEL-CASTRO1,2, Ph. D.
1 Grupo de Investigación en Evolución, Sistemática y Ecología Molecular. INTROPIC. Universidad del Magdalena. Carrera 32# 22 - 08. Santa Marta, Colombia. julianquintero1924@gmail.com. 2 Programa Biología. Universidad del Magdalena. Laboratorio 2. Carrera 32 # 22-08. Sector San Pedro Alejandrino. Santa Marta, Colombia. Tel.: (57 5) 430 12 92, ext. 273.lydaraquelcastro@hotmail.com. Corresponding author: julianquintero1924@gmail.com.
Presentado el 15 de abril de 2013, aceptado el 18 de junio de 2013, correcciones el 26 de junio de 2013.
ABSTRACT
The family Neritidae has representatives in tropical and subtropical regions that occur in a variety of environments, and its known fossil record dates back to the late Cretaceous. However there have been few studies of molecular phylogeny in this family. We performed a phylogenetic reconstruction of the family Neritidae using the COI (722 bp) and the 16S rRNA (559 bp) regions of the mitochondrial genome. Neighbor-joining, maximum parsimony and Bayesian inference were performed. The best phylogenetic reconstruction was obtained using the COI region, and we consider it an appropriate marker for phylogenetic studies within the group. Consensus analysis (COI +16S rRNA) generally obtained the same tree topologies and confirmed that the genus Nerita is monophyletic. The consensus analysis using parsimony recovered a monophyletic group consisting of the genera Neritina, Septaria, Theodoxus, Puperita, and Clithon, while in the Bayesian analyses Theodoxus is separated from the other genera. The phylogenetic status of the species from the genus Nerita from the Colombian Caribbean generated in this study was consistent with that reported for the genus in previous studies. In the resulting consensus tree obtained using maximum parsimony, we included information on habitat type for each species, to map the evolution by habitat. Species of the family Neritidae possibly have their origin in marine environments, which is consistent with conclusions from previous reports based on anatomical studies.
Keywords: Colombian Caribbean, genes mitocondrial, mDNA, Nerita, Neritina, radiation.
RESUMEN
La familia Neritidae cuenta con representantes en regiones tropicales y subtropicales adaptadas a diferentes ambientes, con un registro fósil que data para finales del Cretáceo. Sin embargo no se han realizado estudios de filogenia molecular en la familia. En este estudio se realizó una reconstrucción filogenética de la familia Neritidae utilizando las regiones COI (722 pb) y 16S rRNA (559 pb) del genoma mitocondrial. Se realizaron análisis de distancias de Neighbor-Joining, Máxima Parsimonia e Inferencia Bayesiana. La mejor reconstrucción filogenética fue mediante la región COI, considerándola un marcador apropiado para realizar estudios filogenéticos dentro del grupo. El consenso de las relaciones filogenéticas (COI+16S rRNA) permitió confirmar que el género Nerita es monofilético. El consenso del análisis de parsimonia reveló un grupo monofilético formado por los géneros Neritina, Septaria, Theodoxus, Puperita y Clithon, mientras que en el análisis bayesiano Theodoxus se encuentra separado de los otros géneros. El resultado en las especies del género Nerita del Caribe colombiano fue consistente con lo reportado para el género en estudios previos. En el árbol resultante del análisis de parsimonia se sobrepuso la información del hábitat de cada especie, para mapear la evolución por hábitat. Se obtuvo como resultado que las especies de la familia Neritidae posiblemente tengan su origen en un ambiente marino, siendo congruente con lo reportado en estudios anatómicos realizados anteriormente.
Palabras clave: Caribe colombiano, genoma mitocondrial, mADN, Nerita, Neritina, radiación.
INTRODUCTION
The Neritimorpha (Neritopsina) comprises more than 450 extant species, with a fossil record reported from the Middle Devonian ca 375 million years ago, but possibly as early as Ordovician (Kano et al., 2002). The families Neritidae, Phenacolepadidae, Neritopsidae, Helicinidae, Ceresidae, Proserpinidae, Hydrocenidae, and Titiscaniidae are included in this group (Thompson, 1980; Ponder and Lindberg, 1997; Ponder, 1998).
Among gastropods, Neritimorpha has had one of the greatest adaptive radiation processes. The group has invaded marine, fresh water, and groundwater environments, and exhibits a great variety of forms (Kano et al., 2002). Snails with spiral (several families) or conical forms (Hydrocenidae), with or without opercula, and even slugs that do not develop shells (Titiscaniidae) are included in this group. Some species can be found in terrestrial environments such as those belonging to the families Helicinidae, Ceresidae, Proserpinidae, and Hydrocenidae, whereas other species, namely those in Neritidae, can be found in freshwater and estuarine environments (Thompson, 1980; Ponder, 1998).
The Neritidae has representatives in tropical and subtropical regions, adapted to different environments, and exhibits morphological modifications in various habitats (Holthuis,1995; Kano et al., 2002). This family seems to have its origins in the sea (Kano et al., 2006). About 100 species of the genus Nerita live on marine and intertidal rocks. Species of the genus Smaragdia are found in seagrass areas. However, a higher diversity of Neritidae occurs in freshwater and estuarine waters, in terms of both numbers of genera and of species. Worldwide, 200 species comprise the genera Neritodryas, Clithon, Vittina, Neritina, Neripteron and Septaria (Kano et al., 2002; Kano et al., 2006). Members of the family Neritidae are relatively well represented in the fossil record, dating from the end of the Cretaceous (Bandel, 2008; Frey and Vermeij, 2008).
The evolutionary relations within the family Neritidae have not been well studied, although important studies have been conducted on various genera. For the genus Nerita, there is a very complete analysis of the molecular phylogeny and biogeography of the group in the tropics, using the COI and 16S rRNA genes of the mitochondrial genome and the ATPS subunit of the nuclear genome (Frey and Vermeij, 2008; Frey,2010a). Other studies reconstructed the evolutionary history of the genus Theodoxus and its distribution across the Tethys Sea, using COI and 16S rRNA genes (Bunje and Lindberg, 2007). Studies have also been conducted on the genus Neritina using the COI gene, to analyze the phylogenetic distribution of different reproductive strategies (Kano, 2009). Using anatomical data of species in different genera within Neritopsina, Holthuis (1995), performed a phylogenetic reconstruction of the group, and proposed a phylogeny based on 57 morphological characters. Other studies have used species of the family Neritidae to resolve the phylogeny of Neritimorpha (Neritopsina) and the evolutionary history of the group using nuclear and mitochondrial markers (Kano et al., 2002; Aktipis and Giribet, 2010; Castro and Colgan, 2010).In this study we performed a phylogenetic analysis of the family Neritidae using COI and 16S rRNA regions of the mitochondrial genome, and included species from the Colombian Caribbean. Additionally we reconstructed the evolution of the family using habitat types.
MATERIALS AND METHODS
Study Area
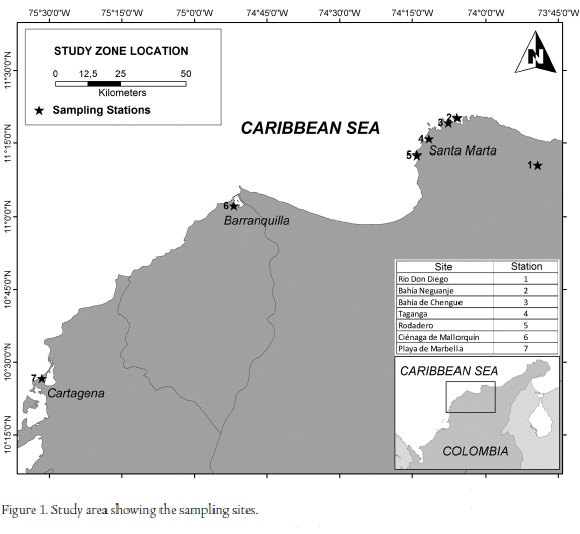
The Colombian Caribbean is located in the northwestern corner of South America (Fig. 1), and includes a coastline of 1937 km, a land area of 7037 km2, and territorial waters of 532162 km2 (Posada et al., 2010). A great variety of environments is represented, including estuarine, marine, and freshwater ecosystems, and the region exhibits a high diversity of organisms.
Collection and Identification of Samples and Sequences
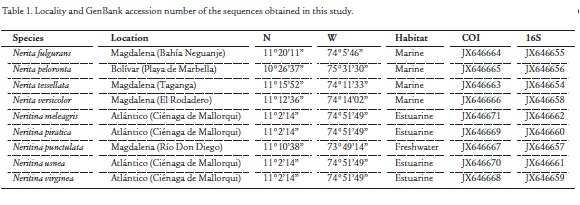
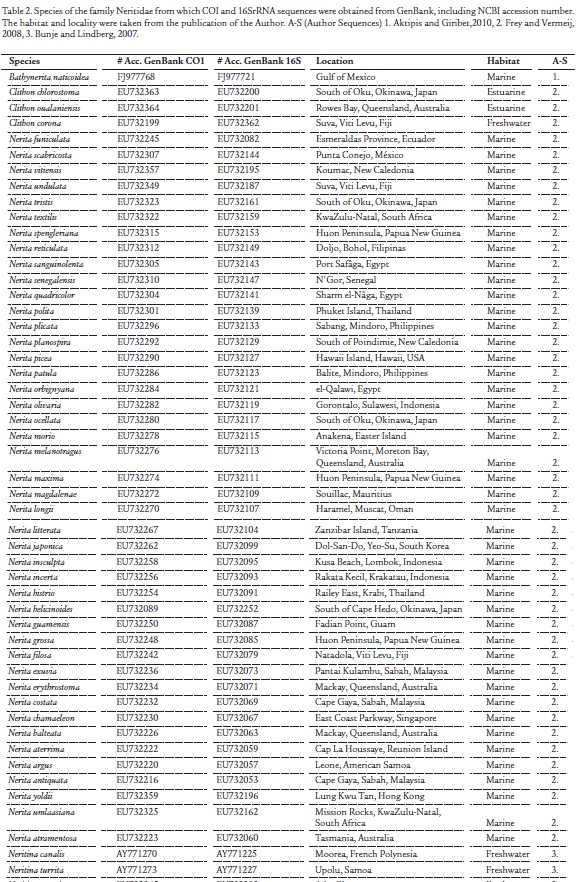
Nine species of neritid snails were collected from different habitats of the Colombian Caribbean (Table 1) and preserved in 96 % ethanol. Samples were identified using morphological taxonomic keys and catalogs (Russell, 1941; Díaz and Puyana, 1994; Yidi and Sarmiento, 2010). Sequences of the COI and 16S rRNA (16S) regions of additional species of the family Neritidae were obtained from GenBank, and were stored with the program MEGA 5 (Tamura et al., 2011), and used in conjunction with the species collected and analyzed from the Colombian Caribbean region. Table 2 shows accession numbers, locality, and habitat for each species downloaded from GenBank. Habitat and locality information were complemented by a literature review. The following total numbers of species by genus were obtained: Puperita (1), Clithon (3), Nerita (44), Neritina (4), Septaria (2) and Theodoxus (3).
DNA Extraction, Amplification and Sequencing
DNA was extracted from the tissue of the foot of each species, using the DNA easy tissue extraction kit (QIAGEN, Valencia, California). The cytochrome oxidase I and 16S rRNA regions of the mitochondrial genome were amplified for each species. PCRs were performed on a total volume of 25 µl. Reactions contained 2,5 µl of 10X buffer, 1 µl of MgCl2 (25 mM), 0,5 µl dNTPs (1 mM), 0,5 µl of Taq polymerase (5 U/mL) (BIOLINE), 2 µl of each primer (10 mM) and 1 µl of DNA.
Amplifications were performed in an Eppendorf gradient PCR thermocycler with the following primer combinations: [COI (HCO2198 5' - 3' TAAACTTCAGGGTGACCAAAAAATCA) and (LCO1490 5' - 3' GGTCAACAAATCATAAAGATATTGG) (Folmer et al., 1994)]; [16S (16Sar 5' - 3' CGCCTGTTTATCAA AAACAT) and (16Sbr 5' - 3' CCGGTCTGAACTCAGATCACGT) (Palumbi, 1996)]. PCR conditions for each gene varied, but generally the amplification consisted of denaturation at 95 °C for 1:00 min, 35 cycles of denaturation at 95 °C 00:15 s, annealing for the COI gene was 46-51 °C, and for the 16S gen was 51-57 °C for 1:00 min, and 1:30 min extension at 72 °C, followed by 5:00 min final extension at 72 °C. PCR optimization for each template involved the variation of MgCl2 concentration and annealing temperature. To remove unincorporated primers and dNTPs before sequencing, double-stranded PCR products were purified using the nucleic acids purification kit of MACHEREY-N. Both strands of the PCR product were sequenced. Primer sequences were removed from the start and the end of the obtained sequence and sequence ambiguities were resolved by comparing the electropherograms using the program BioEdit v. 7.0.5.3 (Hall, 1999). The sequences were submitted to GenBank and are available under Accession numbers JX646654 to JX646671 (Table 1).
Sequence Alignment
CLUSTAL X (Thompson et al., 1997) was used to align the edited sequences and the sequences of the species of Neritidae obtained from GenBank. Representatives of six different genera out of the 16 genera reported for the family were thus included. Alignments were performed using MEGA (v.5) (Tamura et al., 2011). We used MEGA (v.5)to align the COI gene, because this approach can translate the protein-coding nucleotide sequences using the invertebrate mitochondrial genetic code, align the resulting amino acid sequences using Clustal, and then create a nucleotide sequence alignment using the amino acid alignment as a guide. The Clustal settings for the COI gene were: pairwise alignment parameters: gap open penalty = 10, extension penalty = 0,1; multiple alignment parameters: gap open penalty = 10, extension penaly = 0,2; protein weight matrix = Gonnet 250; residue specific penalties = on; hydrophobic penalties = on; gap separation distance = 4; end gap separation = off; negative matrix = off; delay divergent cut-off = 30 %. The Clustal settings for the 16S RNA gene were: pairwise alignment parameters: gap open penalty = 15, extension penalty = 6,66; the multiple alignment parameters were: gap opening penalty = 15, extension penalty = 6,66; DNA weight matrix = IUB; transition weight = 0,5; negative matrix = off; delay divergent cut-off = 30 %.
Sequence Characterization
We analyzed the degree of saturation for the COI gene using the software DAMBE v. 5.3.0 (Xia and Xie, 2001). The percentage of A, T, C and G, together with the percentage of A+ T and G + C for each region was calculated using MEGA (v.5) (Tamura et al., 2011). We also calculated the number of synonymous vs nonsynonymous substitutions for the COI gene (Nei and Gojoborit, 1986) using the model NeiGojobori (Jukes-Cantor). The variance was estimated by the method of bootstraps using 1000 replicates in MEGA (v.5).
Phylogenetic Analysis
For phylogenetic analyses we used three matrices: 1) 16S gene; 2) COI gene; 3) Concatenated COI and 16S genes. Phylogenetic analyses were performed using Neighbor-joining (NJ), maximum parsimony (MP) and Bayesian inference methods (BI). The NJ and MP analyses were conducted in PAUP * version 4.0b10 (Swofford, 2002). Non-parametric bootstrapping was performed using a full heuristic search with 1000 replicates.
Bayesian inference analyses were conducted using Mr.Bayes v. 3.1.2 (Ronquist and Huelsenbeck, 2003). The analysis model was chosen using MrModelTest (Nylander, 2004), with the AIC criterion. The GTR + G + I model was selected for both genes. For each analysis, four chains (three heated, one cold) were run simultaneously for the Monte Carlo Markov Chain. Two independent runs of 15 x 106 generations were performed, with trees sampled every 1000 generations. Each run started from a random tree. Asymptotic convergence to the posterior probability distribution was assessed by examining the plot of generation against the likelihood scores and confirmed using the sump command in software Mr. Bayes. Trees sampled prior to convergence were discarded before construction of the majority rule consensus tree. The percentage of sampled trees recovering a particular clade was used as a measure of that clade s posterior probability (Huelsenbeck and Ronquist, 2001).
Although initial analyses using Haliotis rubra (Vetigastropoda: Haliotidae) and Lophiotoma cerithiformis (Caenogastropoda: Turridae) as outgroups were run, in these cases the internal groups clustered with the outgroup and several inconsistencies in the tree were obtained, consequently we decided to use the species Bathynerita naticoidea as outgroup. B. naticoidea is endemic to the Gulf of Mexico and lives in water depths from 400 m to 2100 m (Zande and Carney, 2001). This species is currently included in the family Neritidae, but it is probably more related to the family Phenacolepadidae, according to evidence from anatomical studies and embryology (Holthuis, 1995; Kano, 2006; Kano et al., 2002). Additionally, for the COI and the concatenated analyses, a BI analysis was performed separating each codon position (1st, 2nd, 3rd) of the COI gene as a partition. The model for each partition was calculated using MrModeltest 2.3, the model assigned to the first position was GTR + G, to the se- cond position was F81 and to the third position was GTR + G. Substitution models and rates of substitution were allowed to vary among the parameters ( unlink command and ratepr = variable ).
Habitat Evolution
We manually mapped the habitat information for each species onto the consensus tree, assigning a different color to each habitat.
RESULTS
Sequence Characterization
After the exclusion of regions of questionable alignment, the concatenated dataset consisted of 1281 characters (722 bp for COI and 559 bp for 16S). For COI, the average frequency of each base was 38.8 % T, 17.5 % C, 22 % A, and 21.7 % G, and the percentage of A+T and G+C was 60.8 % and 39.2 % respectively. For 16S, the average frequency of each base was32.5 % T, 19.6 % C, 29.3 % A, and 18.5 % G, and the percentage of A+T and G+C was 61.9 % and 38.1 % respectively. The number of synonymous substitutions per site for the COI region corresponded to 1,2 ± 0.08 (dS ± SE) and the number of nonsynonymous substitutions per site was 0.06 ± 0.006 (dN ± SE). The dN / dS ratio that defines the type of evolution was equal to 0.054, indicating a negative selection pressure acting on this gene (Pybus and Shapiro, 2009). Some degree of saturation was found for the third codon position of the COI region, Iss 1.9385 > 0.8011 Iss.c, but the position was included in all the analyses because we considered that this site presents valuable phylogenetic information that may not be detected with the first and second codon positions alone (Källersjö et al., 1999; Frey and Vermeij, 2008; Xia, 2009).
Phylogenetic Analysis
Phylogenetic reconstructions of the COI region using Neighbor-Joining (NJ), Maximum Parsimony (MP), and Bayesian (BI) analyses produced similar topologies. The result of the Bayesian analysis for COI (not shown) included a highly supported monophyletic group consisting of species of the genus Nerita; however, inter-node lengths within the genus were very short and in some groups it was not easy to see their internal relationships. The species of the genera Neritina, Septaria, Clithon, and Puperita formed another monophyletic group, in which Neritina virginea presented the longest branch, indicating slightly higher nucleotide substitution rates for this species. The genus Theodoxus formed a monophyletic group separate from the other groups. Although the MP analysis produced a tree with similar topology, the branches were longer and there was better resolution.
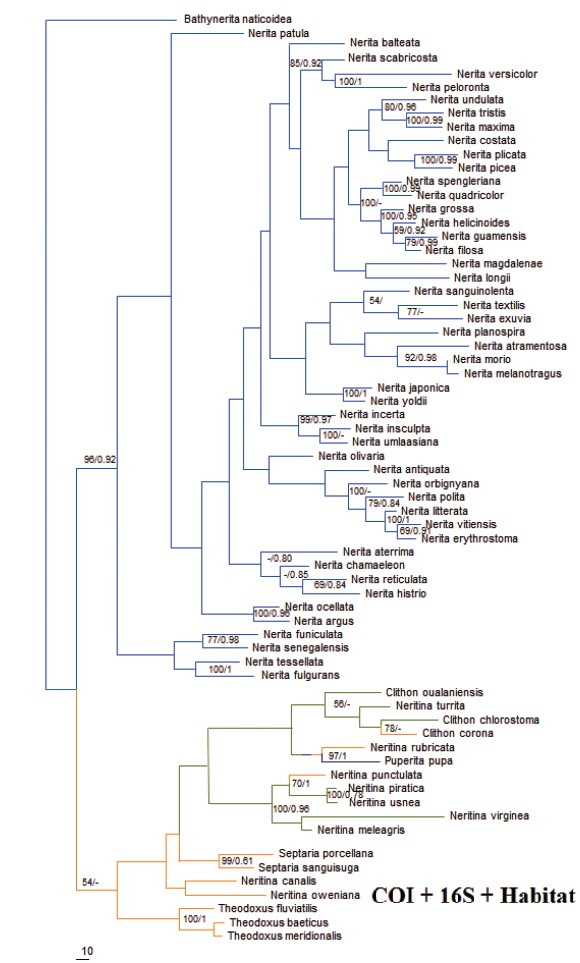
The analysis using the 16S region showed a divergent topology. It showed two groups, one group contained the species of the genus Nerita but with the inclusion of some species of the genus Neritina. In this case, Nerita was not monophyletic. The species of the genera Septaria, Theodoxus, Clithon, and Puperita appeared as a single group, but without a clear divergence and low support values on its nodes (not shown). The consensus phylogenetic reconstructions (COI +16S) using BI formed three groups (Fig. 2a), a monophyletic group formed by the species of the genus Theodoxus, another monophyletic group formed by the species of the genus Nerita and another group with the remaining genera. The monophyly of the genus Nerita and the genus Theodoxus was supported with values of 1.00 and 0.89, respectively (Fig. 2a). In the consensus analyses using MP and NJ, two groups were resolved. A monophyletic group consisting of species of the genus Nerita, with a high bootstrap support of 96 %, and another group with species of the genera Neritina, Puperita, Theodoxus, Septaria, and Clithon with a bootstrap support of 56 % (Fig. 2b). The topology of the MP tree showed longer branches in comparison to the Bayesian analysis, giving better structure to the tree. The partitioned analysis of the consensus dataset (COI 1st + COI 2nd + COI 3rd + 16S) gave the same topology as the non partitioned analysis, however, the analysis without partitions showed higher support values on the nodes.
Mapping Habitat Information
The consensus tree obtained using the parsimony method was used to map the habitat information of each species (Fig. 3). Based on the tree topology, this analysis supports the hypothesis that the family Neritidae originated in marine environments.

DISCUSSION
Phylogeny Of The Family Neritidae
Six percent of the world's known species of Nerita occur in the Colombian Caribbean (Frey and Vermeij, 2008; Frey, 2010b), as do 0.5 % of the species of Neritina that inhabit freshwater and estuarine habitats (Kano et al., 2006). In addition, one species of each of the genera Puperita and Smaragdia, occur in the Colombian Caribbean. As of July 2012, 44 species of the genus Nerita were represented in GenBank, which corresponds to approximately 87 % of the species reported in the world. In contrast, only 13 species belonging to the genera Neritina, Puperita, Septaria, and Clithon were reported in GenBank, representing only 7.5 % of the known species of these groups. Most of these sequences were generated through studies in Europe, Japan and the coasts of North America (Bunje and Lindberg, 2007; Hurtado et al., 2007; Frey and Vermeij, 2008; Kano, 2009; Aktipis andGiribet, 2010). No species of Neritidae known from the Colombian Caribbean were previously represented in Gen Bank. Further, only one complete mitochondrial genome is available for the Neritimorpha, that of the Australasian Nerita melanotragus (Castro and Colgan, 2010). Thus, additional sampling with expanded taxonomic, as well as geographic coverage is needed to further resolve the phylogeny of the family. Specifically, additional studies including poorly represented (Neritina, Puperita, Clithon, Septaria), or unrepresented (i.e., Fluvinerita, Neripteron, Nereina, Clypeolum, Neritodryas) genera are needed. In this study, we sequenced nine species of neritid snails belonging to the genera Nerita and Neritina from the Colombian Caribbean, and used them, together with the sequences available in GenBank to perform an integrated analysis of the family Neritidae.
Our phylogenetic reconstructions revealed a similar topology for COI both with parsimony or BI, separating three groups, one consisting of the monophyletic genus Nerita, another with the species of the genus Theodoxus, and a third group including all the other genera. The analyses showed high support on the basal nodes, confirming that the COI region is a good marker for evaluating and resolving hypotheses about the evolution of the group (Remigio and Hebert, 2003; Frey and Vermeij, 2008). The 16S gene, on the other hand, is considered a good phylogenetic marker in terrestrial mollusks (Klussmann-Kolb et al., 2008), however, the trees produced in our analyses were not consistent, giving different topologies for the MP, NJ and BI analyses. This finding corroborates the results of Frey and Vermeij (2008), who considered the 16S gene as unstable and less useful for reconstructing phylogeny. In our consensus analysis (COI +16S) of the family Neritidae, the genus Nerita is monophyletic. This finding is consistent with the results obtained by Frey and Vermeij (2008) who evaluated the molecular phylogeny and biogeography of Nerita. In our analysis the species of the genera Neritina, Puperita, Septaria, Theodoxus, and Clithon formed a monophyletic group by the parsimony method, however the genus Theodoxus was recovered as a third independent monophyletic group using BI, causing uncertainty about the phylogenetic position of this group within the family, since in both cases the support values were high.
Holthuis (1995), proposed a phylogenetic tree from an anatomical study of the genera and subgenera of the families Neritidae, Phenacolepadidae, and Septariidae, based on 57morphological characters. She considered each genus within the Neritidae as a monophyletic group, an assumption not consistent with the results obtained in the present study, which is the first attempt at integrating all the molecular information available. We strongly recommend the inclusion of a larger number of genera and species, as well as other molecular markers, in studies to further resolve the systematics of the group. Regarding the species collected in the Colombian Caribbean, Nerita versicolor and Nerita peloronta formed a clade that is strongly supported by posterior probabilities and bootstrap values, this clade was always associated with the species Nerita scabricosta. This finding is consistent with what was reported by Frey and Vermeij (2008), who placed these species in the subgenus Nerita sensu stricto. Nerita fulgurans and Nerita tessellata also formed a well supported monophyletic group, which, along with Nerita senegalensis and Nerita funiculata, represents the subgenus Theliostyla (Frey and Vermeij, 2008; Frey, 2010b). The species Neritina punctulata, Neritina piratica, Neritina usnea, Neritina virginea, and Neritina meleagris, appeared as a monophyletic group in the consensus analysis, but the individual analysis of the COI gene recovered N. punctulata + Puperita pupa and N. rubricata + (N. virginea + N. meleagris) as monophyletic groups, and the species N. piratica and N. usnea formed another well supported monophyletic group with species of the genera Clithon and Septaria.
It is important to highlight the close relationship of the species N. piratica and N. usnea that was recovered in all the analyses, which is consistent with Russell's (1941) suggestion that they should be considered a single species (based on conchological characteristics). Since no anatomical or molecular analyses have been used to resolve their taxonomic status, these taxa have continued to be considered as separate species (Yidi and Sarmiento, 2010). The sequences generated in this study for N. piratica and N. usnea, showed no differences in the COI region, and only a three base pair difference in the 16S region between these two species. In contrast, comparison of these sequences with those of N. virginea, showed 19 bp and 123 bp differences for the COI and the 16S regions respectively; N. versicolor and N. peloronta (another very closely related species pair) presented 460 and 257 bp different in the COI and 16S genes, respectively. We conclude that there is little difference at the molecular level between N. piratica and N.usnea, and question their classification as different species. We suggest a more detailed population study and additional examination both at the anatomical and the molecular levels to further assess the status of these taxa.
Evolution of the Family in Relation to Habitat
Adaptive radiation is a response to natural selection and ecological opportunity that involves diversification of species with accompanying adaptations (Glor, 2010). Neritimorpha is a superorder that has undergone significant adaptive radiation, and has an extensive fossil record. It includes representatives that have invaded from marine environments to terrestrial habitats, mainly during the Carboniferous period. Families that currently occur in terrestrial environments include Hydrocenidae, Helicinidae, Proserpinidae, and Ceresidae (Thompson, 1980; Kano et al., 2002; Kano et al., 2006), whereas the family Neritidae includes extant representatives in marine, estuarine, and freshwater environments (Kano et al., 2002; Kano et al., 2006).
Mapping habitat on the phylogenetic reconstruction of the species of the family Neritidae (Fig. 3), makes evident the relationship of the species of the family with marine environments, and strongly suggests evolution from marine to freshwater environments, as proposed by Kano et al., (2006). The genus Nerita, fully occurring in marine habitats, is recovered as a monophyletic group, whereas the species with freshwater and estuarine habitats formed a separate group (although Puperita pupa, which is marine, was recovered within this group). Holthuis (1995) proposed, by an anatomical analysis of the species of the family Neritidae, a parsimonious reconstruction indicating that at least 12 changes have occurred during the evolution among marine, freshwater, and estuarine environments. According to this author, multiple invasions from marine to freshwater environments have occurred. Multiple invasions have also been hypothesized for the genus Septaria entering freshwater streams in tropical Pacific islands (Ponder 1998), and in the radiation of the genus Theodoxus in the river systems of Europe and Central Asia (Bunje and Lindberg, 2007; Bunje, 2007). Some freshwater species of the genus Neritina still have a larval stage in estuarine or marine environments before returning to freshwater streams and rivers (Blanco and Scatena, 2006; Kano, 2009; Gorbach et al., 2012), further supporting the hypothesis that the family has its origins in the sea.
ACKNOWLEDGEMENTS
Thanks to Francisco Borrero, Don Colgan and Tim Pearce for useful suggestions on the preliminary version of the manuscript. This work is part of the products generated in the project 1117-489-25505 code 343-2009 funded by COLCIENCIAS and the University of Magdalena. This work was possible thanks to the scientific research and biodiversity collecting permit No. 15 from the 16 of December 2012 given by ANLA, and the Access to Genetic Resources without commercial interest contract No. 71 between Ministerio de Ambiente y Desarrollo Sostenible and Lyda Raquel Castro.
BIBLIOGRAPHY
Aktipis SW, Giribet G. A phylogeny of Vetigastropoda and other "archaeogastropods": re-organizing old gastropod clades. Invertebr Biol. 2010;129(3):220-240.
Bandel K. Operculum shape and construction of some fossil Neritimorpha (Gastropoda) compared to those of modern species of the subclass. Vita Malacologica.2008;7:19-36.
Blanco JF, Scatena FN. Hierarchical contribution of river - ocean connectivity, water chemistry, hydraulics, and substrate to the distribution of diadromous snails in Puerto Rican streams. JN Am Benthol Soc. 2006;25(1):82-98.
Bunje PME. Fluvial range expansion , allopatry, and parallel evolution in a Danubian snail lineage (Neritidae: Theodoxus). Biol J Linn Soc. 2007;90(4):603-617.
Bunje PME, Lindberg DR. Lineage divergence of a freshwater snail clade associated with post-Tethys marine basin development. Mol Phylogenet Evol. 2007;42(2):373-87.
Castro LR, Colgan DJ. The phylogenetic position of Neritimorpha based on the mitochondrial genome of Nerita melanotragus (Mollusca: Gastropoda). Mol Phylogenet Evol. 2010;57 (2):918-23.
Díaz JM, Puyana M. Familia Neritidae. In: Díaz JM, Puyana M. Moluscos del Caribe Colombiano. INVEMAR, COCIENCIAS, Fundación Natura Colombia. Santa Fe de Bogotá; 1994. p. 122-124.
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotech. 1994;3(5):294-299.
Frey M. A revised classification of the gastropod genus Nerita. The Veliger. 2010a;51(1):1-7.
Frey M. The relative importance of geography and ecology in species diversification evidence from a tropical marine intertidal snail Nerita. J Biogeogr. 2010b;37(8):1515-1528.
Frey M, Vermeij GJ. Molecular phylogenies and historicalbiogeography of a circumtropical group of gastropods (Genus: Nerita): implications for regional diversity patterns in the marine tropics. Mol Phylogenet Evol. 2008;48(3):1067-1086.
Glor RE. Phylogenetic insights on adaptive radiation. Annu.Rev. Ecol. Evol. Syst. 2010;41:251-270.
Gorbach KR, Benbow ME, Mcintosh MD, Burky AJ. Dispersal and upstream migration of an amphidromous neritid snail: implications for restoring migratory pathways in tropical streams. Freshwater Biol. 2012;57(8):1643-1657.
Hall TA. BioEdit: a user-friendly biological sequences aligment editor and analysis program for Window 95/98/NT. Nucl Acid S. 1999;41:95-98.
Holthuis BV. Evolution between marine and freshwater habitats: a case study of the gastropod Neritopsina [PhD thesis]. University of Washington, DC; 1995. p. 249.
Huelsenbeck JP, Ronquist F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics. 2001;17(8):754-755.
Hurtado LA, Frey M, Gaube P, Pfeiler E, Markow TA. Geographical subdivision, demographic history and geneflow in two sympatric species of intertidal snails, Nerita scabricosta and Nerita funiculata, from the tropical eastern Pacific. Mar Biol. 2007;151(5):1863-1873.
Källersjö M, Albert VA, Farris JS. Homoplasy increases phylogenetic structure. Cladistics. 1999;15 (1):91-93.
Kano Y. Usefulness of the opercular nucleus for inferring early development in Neritimorph Gastropods. J Morphol. 2006;267(9):1120-1136.
Kano Y. Hitchhiking behaviour in the obligatory upstream migration of amphidromous snails. Biol Letters. 2009; 5(4):465-468.
Kano Y, Chiba S, Kase T. Major adaptive radiation in Neritopsine gastropods estimated from 28S rRNA sequences and fossil records. T Roy Soc. 2002;269(1508):2457-2465.
Kano Y, Strong EE, Fontaine B, Gargominy O, Glaubrech M, Bouchet P. Focus on freshwater snail. In: Bouchet H, Guyader H, Pascal O. The natural history of Santo. Paris; 2006. p. 257-264.
Klussmann-Kolb A, Dinapoli A, Kuhn K, Streit B, Albrecht C. From sea to land and beyond-new insights into the evolution of Euthyneuran Gastropoda (Mollusca). BMC Evol Biol. 2008;8(1):57-73
Nei M, Gojobori T. Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol. 1986;3(5):418-426.
Nylander J. MrModeltest. Program distributed by the author, Evolutionary Biology Centre. Uppsala University. 2004.
Palumbi SR. Nucleic acids, II: the polymerase chain reaction. In: Hill DM, Morizt C, Mable BK. (Eds.). Molecular Systematics. Massachusetts: Sinauer Associates, Sunderland; 1996. p. 205-247.
Ponder WF. Superorder Neritopsina. In: Beesley P, Ross G, Well A (Eds.). Mollusca: the Southern Synthesis. Fauna of Australia. CSIRO Publishing, Melbourne; 1998. p. 693-702.
Ponder WF, Lindberg DR. Towards a phylogeny of Gastropod Molluscs: An analysis using morphological characters. Zool J Linn Soc-Lond. 1997;119(2):83-265.
Posada BO, Rozo D, Bolaños J, Zamora A. In: INVEMAR Informe del Estado de los Ambientes y Recursos Marinos y Costeros en Colombia Año 2009. Serie de Publicaciones Periódicas No. 8. Santa Marta; 2010. p. 25-32.
Pybus O, Shapiro B. Natural selection and adaptation of molecular sequences. In: Lemey P, Salemi M, Vandamme AM (Eds.). The Phylogenetic Handbook. A practical approach to phylogenetic analysis and hypothesis testing. Cambridge. New York, Madrid; 2009. p. 415.
Remigio EA, Hebert PDN. Testing the utility of partial COI sequences for phylogenetic estimates of Gastropod relationships. Mol Phylogenet Evol. 2003;29(3):641-647.
Ronquist F, Huelsenbeck JP. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 2003;19 (12):1572-1574.
Russell HD. The recent mollusk of the familily Neritidae of Western Atlantic. Bull. Mus. Comp. Zool; 1941. p. 347-404.
Swofford D. PAUP* Phylogenetic analysis using parsimony (*and other methods), version 4. Sinauer Associates, Sunderland, MA. 2002.
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood , evolutionary distance , and maximum parsimony methods research resource. Mol Biol Evol. 2011;28(10):2731-2739.
Thompson FG. Proserpinoid land snails and their relationships within the Archaeogastropoda. Malacologia. 1980;20 (1): 1-33.
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997;25(24):4876-4882.
Xia X. Assessing substitution saturation with Dambe. In Lemey P, Salemi M, Vandamme AM (Eds.). The Phylogenetic Handbook. A practical approach to phylogenetic analysis and hypothesis testing. Cambridge. New York, Madrid; 2009. p. 617-630.
Xia X, Xie Z. DAMBE: Software package for data analysis in molecular biology and evolution. J Hered. 2001(4); 92:371-373.
Yidi E, Sarmiento V. Colombian seashells from the Caribbean sea. Piceno, L Informatore; 2010. p. 1-384.
Zande J, Carney R. Population size structure and feeding biology of Bathynerita naticoidea Clarke 1989 (Gastropod: Neritacea) from Gulf of Mexico hydrocarbon seeps. Gulf Mex Sci. 2001;19 (2):107-118.

Este obra está bajo una licencia de Creative Commons Reconocimiento 3.0 Unported.
Referencias
Aktipis SW, Giribet G. A phylogeny of Vetigastropoda and other "archaeogastropods": re-organizing old gastropod clades. Invertebr Biol. 2010;129(3):220-240.
Bandel K. Operculum shape and construction of some fossil Neritimorpha (Gastropoda) compared to those of modern species of the subclass. Vita Malacologica.2008;7:19-36.
Blanco JF, Scatena FN. Hierarchical contribution of river - ocean connectivity, water chemistry, hydraulics, and substrate to the distribution of diadromous snails in Puerto Rican streams. JN Am Benthol Soc. 2006;25(1):82-98.
Bunje PME. Fluvial range expansion , allopatry, and parallel evolution in a Danubian snail lineage (Neritidae: Theodoxus). Biol J Linn Soc. 2007;90(4):603-617.
Bunje PME, Lindberg DR. Lineage divergence of a freshwater snail clade associated with post-Tethys marine basin development. Mol Phylogenet Evol. 2007;42(2):373-87.
Castro LR, Colgan DJ. The phylogenetic position of Neritimorpha based on the mitochondrial genome of Nerita melanotragus (Mollusca: Gastropoda). Mol Phylogenet Evol. 2010;57 (2):918-23.
Díaz JM, Puyana M. Familia Neritidae. In: Díaz JM, Puyana M. Moluscos del Caribe Colombiano. INVEMAR, COCIENCIAS, Fundación Natura Colombia. Santa Fe de Bogotá; 1994. p. 122-124.
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotech. 1994;3(5):294-299.
Frey M. A revised classification of the gastropod genus Nerita. The Veliger. 2010a;51(1):1-7.
Frey M. The relative importance of geography and ecology in species diversification evidence from a tropical marine intertidal snail Nerita. J Biogeogr. 2010b;37(8):1515-1528.
Frey M, Vermeij GJ. Molecular phylogenies and historicalbiogeography of a circumtropical group of gastropods (Genus: Nerita): implications for regional diversity patterns in the marine tropics. Mol Phylogenet Evol. 2008;48(3):1067-1086.
Glor RE. Phylogenetic insights on adaptive radiation. Annu.Rev. Ecol. Evol. Syst. 2010;41:251-270.
Gorbach KR, Benbow ME, Mcintosh MD, Burky AJ. Dispersal and upstream migration of an amphidromous neritid snail: implications for restoring migratory pathways in tropical streams. Freshwater Biol. 2012;57(8):1643-1657.
Hall TA. BioEdit: a user-friendly biological sequences aligment editor and analysis program for Window 95/98/NT. Nucl Acid S. 1999;41:95-98.
Holthuis BV. Evolution between marine and freshwater habitats: a case study of the gastropod Neritopsina [PhD thesis]. University of Washington, DC; 1995. p. 249.
Huelsenbeck JP, Ronquist F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics. 2001;17(8):754-755.
Hurtado LA, Frey M, Gaube P, Pfeiler E, Markow TA. Geographical subdivision, demographic history and geneflow in two sympatric species of intertidal snails, Nerita scabricosta and Nerita funiculata, from the tropical eastern Pacific. Mar Biol. 2007;151(5):1863-1873.
Källersjö M, Albert VA, Farris JS. Homoplasy increases phylogenetic structure. Cladistics. 1999;15 (1):91-93.
Kano Y. Usefulness of the opercular nucleus for inferring early development in Neritimorph Gastropods. J Morphol. 2006;267(9):1120-1136.
Kano Y. Hitchhiking behaviour in the obligatory upstream migration of amphidromous snails. Biol Letters. 2009; 5(4):465-468.
Kano Y, Chiba S, Kase T. Major adaptive radiation in Neritopsine gastropods estimated from 28S rRNA sequences and fossil records. T Roy Soc. 2002;269(1508):2457-2465.
Kano Y, Strong EE, Fontaine B, Gargominy O, Glaubrech M, Bouchet P. Focus on freshwater snail. In: Bouchet H, Guyader H, Pascal O. The natural history of Santo. Paris; 2006. p. 257-264.
Klussmann-Kolb A, Dinapoli A, Kuhn K, Streit B, Albrecht C. From sea to land and beyond-new insights into the evolution of Euthyneuran Gastropoda (Mollusca). BMC Evol Biol. 2008;8(1):57-73
Nei M, Gojobori T. Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol. 1986;3(5):418-426.
Nylander J. MrModeltest. Program distributed by the author, Evolutionary Biology Centre. Uppsala University. 2004.
Palumbi SR. Nucleic acids, II: the polymerase chain reaction. In: Hill DM, Morizt C, Mable BK. (Eds.). Molecular Systematics. Massachusetts: Sinauer Associates, Sunderland; 1996. p. 205-247.
Ponder WF. Superorder Neritopsina. In: Beesley P, Ross G, Well A (Eds.). Mollusca: the Southern Synthesis. Fauna of Australia. CSIRO Publishing, Melbourne; 1998. p. 693-702.
Ponder WF, Lindberg DR. Towards a phylogeny of Gastropod Molluscs: An analysis using morphological characters. Zool J Linn Soc-Lond. 1997;119(2):83-265.
Posada BO, Rozo D, Bolaños J, Zamora A. In: INVEMAR Informe del Estado de los Ambientes y Recursos Marinos y Costeros en Colombia Año 2009. Serie de Publicaciones Periódicas No. 8. Santa Marta; 2010. p. 25-32.
Pybus O, Shapiro B. Natural selection and adaptation of molecular sequences. In: Lemey P, Salemi M, Vandamme AM (Eds.). The Phylogenetic Handbook. A practical approach to phylogenetic analysis and hypothesis testing. Cambridge. New York, Madrid; 2009. p. 415.
Remigio EA, Hebert PDN. Testing the utility of partial COI sequences for phylogenetic estimates of Gastropod relationships. Mol Phylogenet Evol. 2003;29(3):641-647.
Ronquist F, Huelsenbeck JP. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 2003;19 (12):1572-1574.
Russell HD. The recent mollusk of the familily Neritidae of Western Atlantic. Bull. Mus. Comp. Zool; 1941. p. 347-404.
Swofford D. PAUP* Phylogenetic analysis using parsimony (*and other methods), version 4. Sinauer Associates, Sunderland, MA. 2002.
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood , evolutionary distance , and maximum parsimony methods research resource. Mol Biol Evol. 2011;28(10):2731-2739.
Thompson FG. Proserpinoid land snails and their relationships within the Archaeogastropoda. Malacologia. 1980;20 (1): 1-33.
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997;25(24):4876-4882.
Xia X. Assessing substitution saturation with Dambe. In Lemey P, Salemi M, Vandamme AM (Eds.). The Phylogenetic Handbook. A practical approach to phylogenetic analysis and hypothesis testing. Cambridge. New York, Madrid; 2009. p. 617-630.
Xia X, Xie Z. DAMBE: Software package for data analysis in molecular biology and evolution. J Hered. 2001(4); 92:371-373.
Yidi E, Sarmiento V. Colombian seashells from the Caribbean sea. Piceno, L Informatore; 2010. p. 1-384.
Zande J, Carney R. Population size structure and feeding biology of Bathynerita naticoidea Clarke 1989 (Gastropod: Neritacea) from Gulf of Mexico hydrocarbon seeps. Gulf Mex Sci. 2001;19 (2):107-118.
Cómo citar
APA
ACM
ACS
ABNT
Chicago
Harvard
IEEE
MLA
Turabian
Vancouver
Descargar cita
Visitas a la página del resumen del artículo
Descargas
Licencia
Derechos de autor 2013 Acta Biológica Colombiana

Esta obra está bajo una licencia internacional Creative Commons Atribución 4.0.
1. La aceptación de manuscritos por parte de la revista implicará, además de su edición electrónica de acceso abierto bajo licencia Attribution-NonCommercial-ShareAlike 4.0 (CC BY NC SA), la inclusión y difusión del texto completo a través del repositorio institucional de la Universidad Nacional de Colombia y en todas aquellas bases de datos especializadas que el editor considere adecuadas para su indización con miras a incrementar la visibilidad de la revista.
2. Acta Biológica Colombiana permite a los autores archivar, descargar y compartir, la versión final publicada, así como las versiones pre-print y post-print incluyendo un encabezado con la referencia bibliográfica del articulo publicado.
3. Los autores/as podrán adoptar otros acuerdos de licencia no exclusiva de distribución de la versión de la obra publicada (p. ej.: depositarla en un archivo telemático institucional o publicarla en un volumen monográfico) siempre que se indique la publicación inicial en esta revista.
4. Se permite y recomienda a los autores/as difundir su obra a través de Internet (p. ej.: en archivos institucionales, en su página web o en redes sociales cientificas como Academia, Researchgate; Mendelay) lo cual puede producir intercambios interesantes y aumentar las citas de la obra publicada. (Véase El efecto del acceso abierto).