Aislamiento y Caracterización de Hongos Productores de Fitasa a partir de Muestras Ambientales de Antioquia (Colombia)
Isolation and Characterization of Potential Phytase-Producing Fungi from Environmental Samples of Antioquia (Colombia)
Keywords:
Molecular methods, phosphate, functional food, Aspergillus. / Métodos moleculares, fosfato, alimento funcional, Aspergillus. (es)Downloads
Isolation and Characterization of Potential Phytase-Producing Fungi from Environmental Samples of Antioquia (Colombia)
Aislamiento y Caracterización de Hongos Productores de Fitasa a partir de Muestras Ambientales de Antioquia (Colombia)
Maritza Ocampo Betancur1; Luisa Fernanda Patiño Cervantes2; Mauricio Marín Montoya3; Mauricio Salazar Yepes4 y Pablo Andrés Gutiérrez Sánchez5
1 Biological Engineer. Universidad Nacional de Colombia - Sede Medellín - Faculty of Sciences - School of Biosciences. A.A. 3840, Medellín, Colombia <mocampob@unal.edu.co>
2 Biological Engineer. Universidad Nacional de Colombia - Sede Medellín - Faculty of Sciences - School of Biosciences. A.A. 3840, Medellín, Colombia <lfpatinoc122@gmail.com>
3 Associate Professor. Universidad Nacional de Colombia - Sede Medellín - Faculty of Sciences - School of Biosciences. A.A. 3840, Medellín, Colombia <mamarinm@unal.edu.co>
4 Associate Professor. Universidad Nacional de Colombia - Sede Medellín - Faculty of Sciences - School of Biosciences. A.A. 3840, Medellín, Colombia <masalazay@unal.edu.co>
5 Associate Professor. Universidad Nacional de Colombia - Sede Medellín - Faculty of Sciences - School of Biosciences. A.A. 3840, Medellín, Colombia. <paguties@unal.edu.co>
Recibido: Julio 03 de 2011; aceptado: Marzo 15 de 2012.
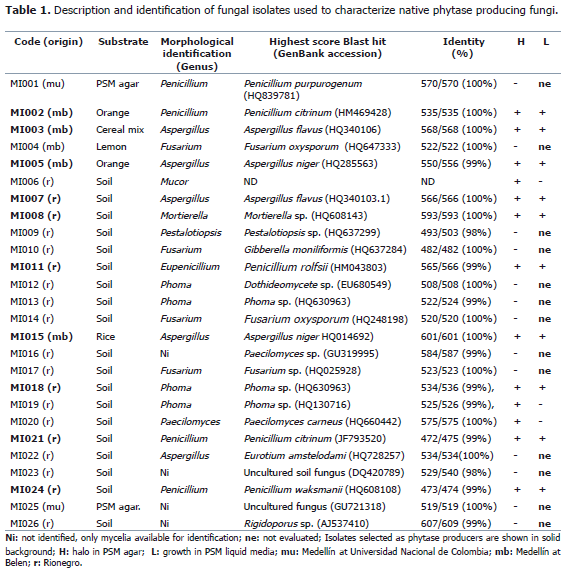
Abstract. Phytases are enzymes used as feed additive that enhance the phosphorus and mineral uptake in monogastric animals and reduce the level of phosphate excretion in their manure. Due to their easy cultivation and high production of extracellular enzymes, filamentous fungi are one of best sources of phytase for use in the feed industry. Phytase has been found principally in the genera Aspergillus, Penicillium, Mucor andRhizopus. In this work, we report the isolation and characterization of environmental fungi producers of phytase with potential use as feed additives. Samples were collected from soils, fruits and cereals in Antioquia (Colombia). A total of 26 fungal strains were isolated and identified using ITS sequencing and morphological analysis. Strains belonged to the following genera: Penicillium, Aspergillus, Fusarium, Mortierella, Pestalotiopsis, Phoma, Paecilomyces and Rigidoporus. Fifty percent of isolates exhibited halos in phytase screening agar indicating that acidic phytases are common enzymes secreted by environmental fungi. Ten isolates were also able to grow in liquid phytase screening medium revealing their potential use for enzyme production in submerged fermentations. Molecular detection of the PhyA gene from Aspergillus was achieved. Partial sequence of the phyA gene from one A. niger isolate was obtained and analyzed.
Key words: Molecular methods, phosphate, functional food, Aspergillus.
Resumen. Las fitasas son enzimas utilizadas como aditivo en productos de alimentación animal, con el fin de mejorar la asimilación de fósforo y minerales en animales monogástricos y disminuir la excreción de fósforo al ambiente. Los hongos filamentosos son una de las mejores fuentes de fitasas debido a su facilidad de cultivo y altos niveles de producción de enzimas extracelulares. Los principales productores de fitasas corresponden a miembros de los géneros Aspergillus, Penicillium, Mucor y Rhizopus. En este trabajo se reporta el aislamiento y caracterización de hongos ambientales productores de fitasas con aplicación potencial en la industria de alimentación animal. Se obtuvieron e identificaron un total de 26 aislamientos; caracterizados por secuenciación de la región ITS-ADNr y análisis morfológico. Los aislamientos pertenecieron a los siguientes géneros:Penicillium, Aspergillus, Fusarium, Mortierella, Pestalotiopsis, Phoma, Paecilomyces y Rigidoporus. Se observó la secreción de fitasas en 50% de los aislamientos sugiriendo la ubiquidad de esta enzima en hongos ambientales. Diez aislamientos crecieron eficientemente en medio líquido con fitato como única fuente de fósforo. Estos últimos cumplen con los requisitos para la producción de enzimas mediante fermentación sumergida. Se diseñaron cebadores para la detección molecular del gen PhyA en los aislamientos del géneroAspergillus. Se obtuvo y analizó la secuencia parcial del gen PhyA de un aislamiento de A. niger.
Palabras clave: Métodos moleculares, fosfato, alimento funcional, Aspergillus.
Phytate (myo-inositol 1,2,3,4,5,6-hexakisphosphate; IP6) is a phosphorylated derivative of myo-inositol important in the storage and retrieval of phosphorus, inositol and ions during plant development and germination (Raboy, 2003). The phosphorus fraction stored as phytate range from 30% in roots and up to 80% in seeds and cereals (Oh et al., 2004). Animal feed is primarily made out of oilseed meals, cereal grains and legumes which contain considerable amounts of phytate. Unfortunately, this phosphorous source is unavailable to monogastric animals like pigs and poultry due to the lack of appropriate enzymes (Vats and Banerjee, 2004). To compensate for phosphorus intake, animal feed is supplemented with inorganic phosphate, an expensive and scarce mineral. Phytate also acts as a chelator of metal ions preventing the normal intake of calcium, magnesium, zinc and iron (Murry et al., 1997). Finally, phosphorus from phytate can end up in manure, giving rise to serious environmental problems such as algal blooms and eutrophication of surface water in areas of intensive farming (Vats and Banerjee, 2004).
Phytases are enzymes used as feed additive that enhance the phosphorus and mineral uptake in monogastric animals and reduces the level of phosphate output in their manure. Degradation of phytate by phytase avoids the necessity to add inorganic phosphate and reduces the amount of phosphorus released to the environment (Haefner et al., 2005). Supplementation of swine and poultry diets with microbial phytases can improve significantly the bioavailability of phytate P and reduce P excretion. When added to pig and poultry feed, the normal addition of phosphate can be lowered considerably, and the amount of undigested phytate in the manure of the animals is reduced up to 30-60% (Vats and Banerjee, 2004). Cromwell et al. (1993) tested the efficacy of a microbial phytase produced by Aspergillus niger in corn-soybean meal or dextrose-corn starch-soybean meal-based diets and were able to prove that phytase is an efficient way of improving the bioavailability of phytate P for pigs. Addition of microbial phytase to diets for growing pigs can increase the apparent absorbability of P by 24% and reduce the amount of phosphate in feces by 35%. More recent studies have shown that addition of phytase improve the protein efficiency ratio of broiler chickens (Kong and Adeola, 2011). Lei et al. (2011) have shown that entire substitution of inorganic phosphorus by phytase can significantly increase bone strength, and Ca and P contents in tibia ash in birds. Phytase supplementation improves digestibility and retention of Ca, P and N in starter and growing pigs without any negative effect on pig performance (Columbus et al., 2010). Transgenic plants that excrete phytases into soil are also being investigated with a view to increasing the inorganic phosphate available for absorption (George et al., 2005).
Phytases are widespread enzymes and can be found in animals, plants, and microorganisms. However, due to their easy cultivation and high production of extracellular enzymes, filamentous fungi are one of best sources of phytase for use in the feed industry. Phytase has been found principally in the genera Aspergillus, Penicillium, Mucor and Rhizopus (Vats and Banerjee, 2004). Other phytase producing strains include Cladosporium species, Myceliophthora thermophila, Emericella nidulans, Talaromyces thermophilus and Thermomyces lanuginosus (Mitchell et al., 1997; Pasamontes et al., 1997). However, several studies have confirmed Aspergillus strains to be the best producers of extracellular phytase (Howson and Davis, 1983; Casey and Walsh, 2003; Vats and Banerjee, 2004; Xiong et al., 2004; Zhang et al., 2010). In this work, we report the isolation and characterization of environmental fungi producers of phytase from Antioquia with potential use as feed additives.
MATERIALS AND METHODS
Isolation of microorganisms. Fungi were isolated using various substrates like soil, grains cereals and fruits as shown in Table 1. A total of 19 fungal strains from soil were isolated using serial dilutions. Briefly, 1 g of soil from the municipality of Rionegro (Antioquia, Colombia) was dissolved in 9 mL of sterile distilled water and dilutions of 10-2, 10-3 and 10-4 were plated onto sterilized potato dextrose agar (300 g potato, 20 g dextrose, 20 g agar per liter) containing 100 mg mL-1 of ampicillin and incubated at 25 °C in the laboratory for one week. Fungi growing on agar plates were subcultured in fresh PDA medium until pure colonies were observed. Seven environmental isolates from various locations in Medellín (Universidad Nacional de Colombia - Sede Medellín and district of Belen) were collected in moist chambers using 250 mL styrofoam cups with a layer of wet filter paper at the bottom. Grains (rice, wheat and a mix of wheat, barley and oat flakes) and fruits (lemon and orange) were used as substrate. These chambers were left open at the site of collection for one day and then closed and incubated at room temperature. Fungal growth was observed within one week of incubation. Isolation and purification was done on PDA plates as described above. A total of 26 fungal isolates were obtained in pure cultures by single spore transfer onto PDA plates and stored at 4 °C in the laboratory of Industrial Microbiology, Universidad Nacional de Colombia - Sede Medellín for further use. Isolates were preserved by immersing a holepunch of mycelium in 20% sterile glycerol and stored in -80 °C freezer. Isolated fungi were preliminarily identified on the basis of morphological characteristics using updated taxonomical keys (Seifert et al., 2011) and MycoBank (Crouset al., 2004).
Phytase screening. Screening was carried out on agar plates containing sodium phytate (4 g L-1) as sole phosphorus source (Howson and Davis, 1983). Ten fungi were selected based on their ability to form halos. Positive results from the screening were confirmed by growing the fungi in PSM medium using submerged fermentation. Liter composition of PSM was: 20 g glucose, 4 g sodium phytate, 2 g CaCl2, 5 g NH4NO3, 0.5 g KCl, 0.5 g MgSO4.7H2O, 0.01 g MnSO4.H2O and 0.01 g FeSO4.7H2O. pH was adjusted to 5.5 by drop wise addition of 0.1M HCl. 30 mL aliquots were autoclaved using a Trident Medical horizontal autoclave, at 15 psi, 120 °C for 20 min, in 100 mL Erlenmeyer flasks, inoculated with a holepunch of mycelium and incubated for up to seven days at 30 °C in an Innova 4400 orbital shaker at 150 rpm. A positive control supplemented with an equivalent amount of sodium phosphate and a negative control lacking phosphorous. Preparation of controls was similar to the PSM media described above.
Molecular characterization. 26 isolates were grown on 2% malt extract (ME; 20 g malt extract in 1 L deionised water) for 12 days at room temperature. Mycelium was collected and macerated in liquid nitrogen. DNA was purified using the DNeasy Plant Mini Kit (Qiagen) using the manufacturers protocol. The ITS1, 5.8S and ITS2 regions of the ribosomal RNA operon were amplified using primers ITS1 (5'-TCC GTA GGT GAA CCT GCG G-3') and ITS4 (5'-TCC TCC GCT TAT TGA TAT GC-3) reported by White et al. (1990). The 50 ml polymerase chain reaction (PCR) mixture included 0.2 mM of each dNTP; 0.4 mM of each primer; 2 mM MgCl2; buffer Fermentas 1X; 2.5 U of Taq polymerase (Fementas) and 5-10 ng of DNA template. PCR amplification consisted of an initial denaturation step at 98 °C for 3 min, followed by 35 cycles of 60 s at 95°C, 60 s at 55 °C and 70 s at 72 °C. Final chain elongation was achieved at 72 °C for 10 min. PCR products were separated in 1.5% agarose gel electrophoresis and then purified using Qiagen PCR purification kit for direct sequencing using an ABI PRISM Big DYE Terminator Cycle Sequencing Ready Reaction Kit (Applied BioSystems, Foster City, CA). Sequencing results were assembled using CONSED (Gordon et al., 1998). Each ITS sequence was used to perform BLASTN searches at the NCBI website (http://www.ncbi.nlm.nih.gov/BLAST/). Preliminary annotation of each isolate was based on the BLAST hit with maximun score to the query sequence. Phylogenetic affinities were determined for fungus considered positive for phytase production by comparison with sequences of related species retrieved from GenBank. Multiple alignments were generated using CLUSTAL (Thompson et al., 1994). Phylogenetic relationships were estimated using MEGA program Version 4.0 (Tamura et al., 2007). Phylogenetic trees were constructed using the Neighbor-Joining (NJ) algorithm with bootstrap values calculated from 1,000 replicate runs. The Maximum Composite Likelihood model was used to estimate evolutionary distance.
Phytase gene amplification. Due to the importance of Aspergillus as a phytase producer, a set of degenerate primers was designed to amplify a fragment of the PhyA gene. Primers were designed based on a multiple alignment of the following sequences (Uniprot accession in parentheses): A. niger (P34752), A. fumigatus(O00092) and A. terreus (O00085 and O00100). Sequences were obtained from UniProt, EXPASY (UniProt Consortium, 2008). Primers were obtained with CODEHOP using the protein alignments as input (Staheli et al., 2011) and were: PhyAspF, 5'-GACACCGTGGACCAGggntaycartg-3' (degeneracy = 16, Tm = 61.0 °C) and PhyAspR, 5' -GCAGATGCCGTGGTCCarngtrttrtt-3' (degeneracy = 32, Tm=63.0 °C). PCR was carried out in a volume of 10 mL with the same reagent concentration used for the amplification of the ITS regions. The PCR program consisted of an initial denaturation step at 98 °C for 3 min, followed by 35 cycles of 60 s at 95°C, 60 s at 41 °C and 60 s at 72 °C. Final chain elongation was achieved at 72 °C for 8 min. PCR products were separated in 1.5% agarose gel electrophoresis and sequenced as described.
RESULTS AND DISCUSSION
Preliminary screening. A total of 26 fungal strains were isolated and identified using ITS sequencing and morphological analysis (Table 1). Colonial morphology of all strains is shown in Figure 1. According to Blast strains belonged to the following genera: Penicillium (5), Aspergillus (4), Fusarium (3), Mortierella (1), Pestalotiopsis (1),Gibberella (1), Phoma (3), Eurotium (1), Paecilomyces (2), and Rigidoporus (1). Sequencing results were not available for isolate MI006. Blast hits for isolates MI023 and MI025 corresponded to uncultured fungus while MI012 was only identified as belonging to class Dothideomycetes. Morphological analysis of each isolate confirmed the Blast identification at the genera level for 17 strains (Table 1). Isolates MI010 and MI022 were identified asFusarium sp. and Aspergillus sp., respectively. For these isolates Blast results coincided with their teliomorphic states: Gibberella and Eurotium, respectively. Isolate MI011 was observed and identified as Eupenicillium, which corresponds to the teliomorph state of Penicillium in the molecular identification. In spite of lacking DNA sequence information for isolate MI006, it was identified morphologically as Mucor sp. Morphological identification of isolates MI016, MI023 and MI026 was not possible as neither spores nor reproductive structures were observed.
The most common method for detecting phytase activity relies on the disappearance of precipitated sodium phytate as an indication of enzyme activity. In this assay, fungi secreting phytases produce clear zones on agar media containing sodium or calcium phytate (Shieh and Ware, 1968; Howson and Davis, 1983). According to this test, 50% of 26 isolates produced extracellular phytase (Table 1). Clear zones were observed for isolates belonging to five different genera: Penicillium (4), Aspergillus (4), Paecilomyces (1), Mortierella (1), Mucor (1), and Phoma (2) (Figure 2). With the exception of MI022, MI023 and MI025, some strains showed significant growth but failed to show clear zones in PSM agar. However, sodium phytate was the only phosphorous source in the media and agarose was used instead of agar to guarantee that trace phosphate was maintained to a minimum. It is possible that some strains use intracellular or/and alkaline phytase, which will not form halos, to utilize this phosphorous source.
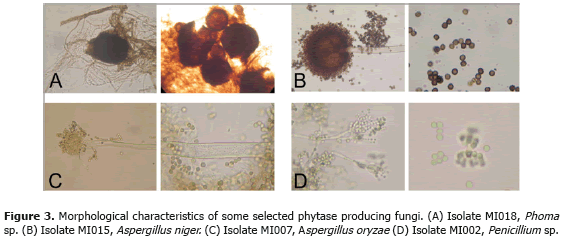
Growth in liquid media. It has been shown that PSM can give some false positives with acid producing strains (Bae et al., 1999). As useful phytase producers should grow efficiently in liquid media, a second screening was performed. Fungal growth in liquid PSM was compared to a negative control lacking any phosphorous source. According to this criterion, strains MI002, MI003, MI005, MI007, MI008, MI011, MI015, MI018, MI021 and MI024 were chosen as phytase producers of interest (Table 1). Phylogenetic analysis confirmed that these strains belonged to the genera Phoma, Mortierella, Penicillium and Aspergillus. Morphological features of selected strains are shown in Figure 3. Phytase producing strains.
Phoma. Isolates MI018 and MI019 formed halos in solid PSM but only MI018 showed significant growth in liquid media. The phylogenetic affinity for these strains is shown in Figure 3A. MI018 is closely related to Phoma herbarum and Phoma macrostoma (Figure 4A). P. herbarum is a ubiquitous saprophyte and versatile producer of many natural products such as gibberellins and haloquinones (Hamayun et al., 2009). P. macrostoma is a poorly studied fungus originally described by Westendorp in 1852 (Crous et al., 2004). Recent reports have shown this species to be a major producer of tyrosol (Tyagunova and Sorensen, 2010). P. macrostoma has also been shown to have bioherbicidal effects on weeds (Bailey and Derby, 2001). Interestingly, this is probably the first report suggesting phytase production by members of this genus
Mortierella. Mortierella isolate (MI008) produced halos in PSM agar together with significant growth in liquid PSM. Determination of the phylogenetic affinity at the species level was not possible due to the absence of well-curated sequences in GenBank. However, it is clear that MI018 belongs to Mortierella and not to the related genus Umbelopsis (Figure 4B). Mortierella is a fungus that belongs to the class Zygomycetes, commonly found in soil, rhizosphere and plants or animals that remains in contact with soil (Domsch et al., 1980; Webster and Weber, 2007). Some species are producers of polyunsaturated lipids of nutritional value such as arachidonic acid and have been used in the production of pharmaceuticals (Dyal and Narine, 2005). Mortierella can be isolated on nutrient-poor media and can grow well at 40-42°C (Austwick, 1976). So far there are no reports of phytases produced by Mortierella. No genes coding for phytase were found in GenBank either. However there exist some reports of phytase production by fungi of the order Mucorales, such as Mucor piriformis (Howson and Davis, 1983) and Rhizopus oligosporus (Casey and Walsh, 2004). It has been shown that rhizosphere isolates ofMortierella sp. can solubilize rock phosphate and increase the P content of mycorrhizal plants (Osorio and Habte, 2001). Similar results were observed by Zhang et al., (2011) who demonstrated that Mortierella sp. is an efficient phosphate solubilizing fungus that can significantly affect plant growth. It would not be surprising that phytase is part of the enzymatic pool of this phosphate solubilizing fungus. Genome sequencing projects of Mortierella alpina ATCC 32222 and M. verticillata are underway (http://www.ncbi.nlm.nih.gov/sites/genome). These projects will give great insight on the phosphate metabolism of these organisms and could confirm phytase production by this soil fungus.
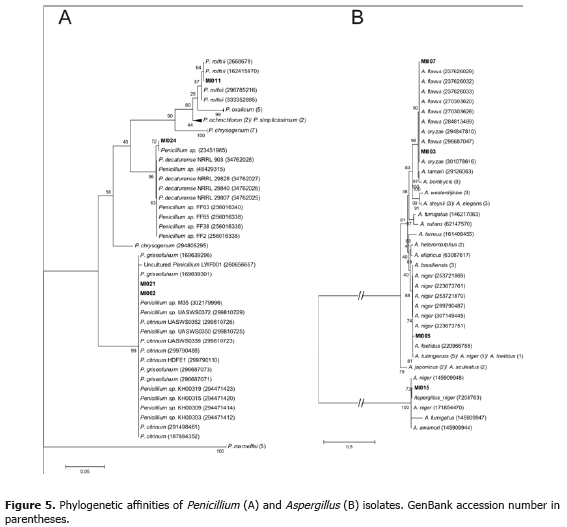
Penicillium. Penicillium has a prominent industrial use due to its ability to produce large quantities of extracellular enzymes (Webster and Weber, 2007). It comes as no surprise that four out of five Penicillium isolates, MI002, MI011, MI021 and MI024, were considered potential phytase producers. Phylogenetic affinities as deduced from NJ analysis are shown in Figure 5A. MI0011 and MI0024 were identified as P. rolfsii and P. decaturense, respectively. Unfortunately, sequence analysis of strains MI002 and MI021 was not able to resolve their affinity to either P. citrinum or P. griseofulvum. Identification at the species levels for isolates MI002 and MI021 will require more detailed analysis. P. rolfsii is a poorly studied species originally described in 1930 by Thom (Crous et al., 2004). There are no current reports on the biotechnological application of this fungus. P. decaturense is a species isolated from wood-decay and recently described by Peterson et al. (2004).
This species is of interest as it can be the source of novel bioactive insecticidal compounds (Zhang et al., 2003).P. citrinum is a common endophytic fungus of cereal plants like wheat and soybean studied for its production of citrinin and secondary metabolites (Wakiyama et al., 2008). P. griseofulvum has been associated with blue mold decay in storage apple fruits (Moslem et al., 2010), production of metabolites with potential activity against cancer cells (Wang et al., 2009) and inhibitors of diacyglycerol acyltransferases (Lee et al., 2008). A search forPenicillium phytases in GenBank retrieved only six results: P. oxalicum (AY071824, HM053476), P. chrysogenum(XM_002561048), P. marneffei (XM_002148785, XM_002147131) and Penicillum sp. Q7 (EF197827). However, none of our isolated species have been studied for phytase production.
The first detailed report on the isolation and characterization of a Penicillium phytase dates back to 2000 (Tsenget al., 2000). After screening eighty-three isolates from different soil samples the most active fungal isolate with phytase activity was identified as P. simplicissimum. The purified enzyme showed an optimum pH and temperature of pH 4.0 and 55 °C, respectively. This phytase revealed broad substrate specificity and was strongly inhibited by Fe2+, Fe3+ and Zn2+ (Tseng et al., 2000). Lee et al. (2007), characterized a novel fungal phytase from P. oxalicum PJ3, cloned its gene and studied the expression in P. pastoris. The pH optimum of the purified enzyme was 4.5 and an optimal temperature of 55 °C. In a more recent work, Zhao et al. (2010) isolated a protease-resistance phytase gene of Penicillium sp. and successfully isolated mutants with improved thermal stability and optimal temperature and pH. The authors suggest that these improved properties make these engineered enzymes more suitable to be used as feed additive in the feed industry than the present commercial phytases.
There are few detailed investigations on the application of Penicillium phytases. Kozlowski et al. (2009), tested the species P. canescens phytase in broiler rations with considerably reduced levels of phosphorus and showed that the process of bone mineralization was enhanced by phytase supplementation. Supplementation of pig diets with Penicillium phytase increased the apparent total tract digestibility of phosphorus and calcium by 22.6 and 18.3 percent, respectively (Steinera et al., 2006). The effect of P. purpurogenum phytase in enhancing the mobilization of native unavailable phosphorus was evaluated in arid environments. A significant improvement in plant biomass (30%), root length (21%), P uptake (6%), seed (19%) and straw yield (30%), and P concentration of shoot (15%), root (6%), and seed (33%) resulted from inoculation of P. purpurogenum (Yadava and Tarafdara, 2011). In spite of the limited studies available on Penicillium phytases, these results suggest interesting applications for this kind of enzymes. It would be interesting to investigate further the biochemical properties of phytases produced by isolates MI002, MI011, MI021 and MI024.
Aspergillus. This genus is the most widely used by industry for the production of enzymes and organic acids (Webster and Weber, 2007). Aspergillus phytases have shown a series of desirable features such as thermal stability, substrate specificity and activity at low pH. For this reason the majority of commercial phytases originate from this group (Vats and Banerjee, 2004). In our study, four out of five Aspergillus isolates showed potential phytase activity: MI003 (A. flavus), MI005 (A. foetidus), MI007 (A. flavus) and MI015 (A. niger). Phylogenetic affinities for the Aspergillus isolates as deduced from NJ analysis are shown in Figure 5B. Strains MI003 and MI007 were classified as A. oryzae instead of A. flavus. However, the former is considered to have resulted from the domestication of A. flavus. A comparison of both genomes confirmed this hypothesis and concluded that these two fungi may be ecotypes of the same domestication of A. flavus. In nature, A. flavus is capable of growing on many nutrient sources. It is predominately a saprophyte and grows on dead plant and animal tissue in the soil. A. flavus can infect seeds of corn, peanuts, cotton, and nut trees (Payne et al., 2006). A. foetidus has been used in the production of enzymes such as xylanase, alpha-galactosidase, amylase and tannase (Michelena and Castillo, 1984; Purohit et al., 2006; Liu et al., 2007; Valte et al., 2010).
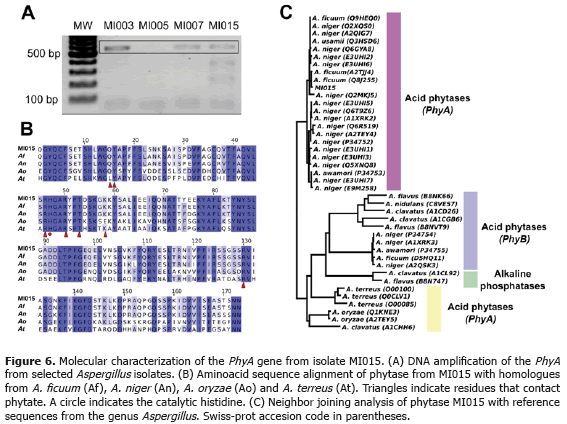
There are several published reports of phytase production by species of the Aspergillus genus: A. niger NRRL 3135, A. flavus, A. terrus, A. carneus, A. oryzae and A. fumigatus (Shieh and Ware, 1968; Shieh et al., 1969; Ohet al., 2004). However, the most widely used phytases are isolated from Aspergillus niger (Xiong et al., 2005), A. fumigatus (Pasamontes et al., 1997) and A. ficcum (Mitchell et al., 1997). A database search in GenBank demonstrated the presence of phytase in A. awamori, A. clavatus NRRL, A. ficuum, A. flavus NRRL3357, A. nigervar awamori, A. niger CBS 513, A. oryzae, Aspergillus sp. A25 A. terreus and A. usamii strain N-2418. Using these sequences, a set of degenerate primers was designed in this study to detect Aspergillus phytases. PCR amplification gave bands with the expected size (~500 bp) for isolates MI003, MI005 and MI015 (Figure 6A). To confirm that this PCR product corresponded to the phytase gene, the 500 bp from MI0015 was sequenced. A Blastx search of the sequenced amplicon revealed that this sequence shared 99% nucleotide identity with PhyAgene from A. ficuum (AF537344.1) and 98% with PhyA from A. niger CBS 513.88 (XM_001401676.2). Multiple alignment of the protein sequence of MI015 phytase revealed its high similarity with A. ficuum, A. niger, A. oryzaeand A. terreus phytases (Figure 6B). It is important to note that A. niger PhyA is the active ingredient of Natuphos™(BASF animal nutrition), the commercial phytase most commonly used as additive in animal feed (Wodzinski and Ullah, 1996).
There are three main types of phytases in Aspergillus: PhyA, PhyB and alkaline phytases. PhyA and PhyB phytase belong to the family of histidine acid phosphatases (HAP) characterized by a RH(G/N)XRXP motif, unique to this enzyme class (Oh et al., 2004). PhyA phytases are active in a wide pH range (2.5-5.0) while PhyB are capable of hydrolyzing phytate quite efficiently at pH 2.5 but have very little activity at pH 5.0 (Ullah and Phillipy, 1994). HAP can hydrolyze metal-free phytate at acidic pH. Alkaline phytases have an optimum pH of 7.0-8.0 and can hidrolyze phytate in complex with metal cations (Oh, et al., 2004). Phylogenetic analysis confirmed the classification of MI015 phytase as a PhyA histidine acid phytase (Figure 6C), a desirable condition for future biotechnological applications.
CONCLUSIONS
To our knowledge, this is the first systematic study aimed at the isolation of phytase producing fungi in Colombia. The present investigation suggests that phytase producing fungi can be readily isolated from soils and grains. Fifty percent of all our isolates exhibited halos in phytase screening agar indicating that acidic phytases are a common enzyme secreted by environmental fungi. Ten isolates were also able to grow in liquid phytase screening medium revealing their potential use for enzyme production in submerged fermentations. These results encourage further studies aimed at the isolation and biochemical characterization of native phytase producing fungi as a preliminary step in their use as feed additives and production at industrial scale.
ACKNOWLEDGEMENTS
This research was supported by a grant from International Foundation for Science (IFS, Sweden; Agreement No. F/4634-1) and DIME (20101007951). We thank Doctor Walter Osorio from the Soil Microbiology Laboratory at Universidad Nacional de Colombia, Sede Medellín and Johana Gutiérrez for their valuable help.
BIBLIOGRAPHY
Austwick, P.C.K. 1976. Environmental aspects of Mortierella wolfii in cattle. New Zealand Journal of Agricultural Research 19(1): 25-33.
Bae, H.D., L.J. Yanke, K.J. Cheng and L.B. Selinger. 1999. A novel staining method for detecting phytase activity. Journal of Microbiological Methods 39(1): 12-22.
Bailey, K.L. and J. Derby. 2001. Fungal isolates and biological control compositions for the control of weeds. US Patent Application Serial No. 60/294,475. Filed 20 May 2001.
Casey, A. and G. Walsh. 2003. Purification and characterization of extracellular phytase from Aspergillus nigerATCC 9142. Bioresource Technology 86(2): 183-188.
Casey, A. and G. Walsh. 2004. Identification and characterization of a phytase of potential commercial interest. Journal of Biotechnolology 110(3): 313-322.
Columbus, D., S.J. Niven, C.L. Zhu and C.F. de Lange. 2010. Phosphorus utilization in starter pigs fed high-moisture corn-based liquid diets steeped with phytase. Journal of Animal Science 88(12): 3964-3976.
Cromwell, G.L., T.S. Stahly, R.D. Coffey, H.J. Monegue and J.H. Randolph. 1993. Efficacy of phytase in improving the bioavailability of phosphorus in soybean meal and corn-soybean meal diets for pigs. Journal of Animal Science 71(7): 1831-1840.
Crous, P.W., W. Gams, J.A. Stalpers, V. Robert and G. Stegehuis. 2004. MycoBank: an online initiative to launch mycology into the 21st century. Studies in Mycology 50: 19-22.
Domsch, K.H., W. Gams and T.H. Anderson. 1980. Compendium of soil fungi. Volume 1. Academic Press, London. 406 p.
Dyal, S.D. and S.S. Narine. 2005. Implications for the use of Mortierella fungi in the industrial production of essential fatty acids. Food Research International 38(4): 445-467.
George, T.S., R.J. Simpson, P.A. Hadobas and A.E. Richardson. 2005. Expression of a fungal phytase gene inNicotiana tabacum improves phosphorus nutrition of plants grown in amended soils. Plant Biotechnology Journal 3(1): 129-140.
Gordon, D., C. Abajian and P. Green. 1998. Consed: a graphical tool for sequence finishing. Genome Research 8(3): 195-202.
Haefner, S., A. Knietsch, E. Scholten, J. Braun, M. Lohscheidt and O. Zelder. 2005. Biotechnological production and applications of phytases. Applied Microbiology and Biotechnology 68(5): 588-97.
Hamayun, M., S.A. Khan, A.L. Khan, G. Rehman, E.Y. Sohn, A.A. Shah, S.K. Kim, G.J. Joo and I.J. Lee. 2009.Phoma herbarum as a new gibberellin-producing and plant growth-promoting fungus. Journal of Microbiology and Biotechnology 19(10): 1244-1249.
Howson, S.J. and R.P. Davis. 1983. Production of phytate hydrolyzing enzyme by some fungi. Enzyme and Microbiology Technology 5(5): 377-389.
Kong, C. and O. Adeola. 2011. Protein utilization and amino acid digestibility of canola meal in response to phytase in broiler chickens. Poultry Science 90(7): 1508-1515.
Koztowski, K., J. Jankowski and H. Jeroch. 2009. Efficacy of different phytase preparations in broiler rations. Polish Journal of Veterinary Sciences 12(3): 389-393.
Lee, J., C. Yunjaie, L. Peter-Changwhan, K. Seungha, B. Jinduck and C. Jaiesoon. 2007. Recombinant production of Penicillium oxalicum PJ3 phytase in Pichia pastoris. World Journal of Microbiology and Biotechnology 23(3): 443-446.
Lee, S.W., M.C. Rho, J.H. Choi, K. Kim, Y.S. Choi, H.S. Lee and Y.K. Kim. 2008. Inhibition of diacylglycerol acyltransferase by phenylpyropenes produced by Penicillium griseofulvum F1959. Journal of Microbiology and Biotechnology 18(11):1785-1788.
Lei, Q.B., L.X. Shi, K.Y. Zhang, X.M. Ding, S.P. Bai and Y.G. Liu. 2011. Effect of reduced energy, protein and entire substitution of inorganic phosphorus by phytase on performance and bone mineralisation of laying hens. British Poultry Science 52(2): 202-13.
Liu, C., H. Ruan, H. Shen, Q. Chen, B. Zhou, Y. Li and G. He. 2007. Optimization of the fermentation medium for alpha-galactosidase production from Aspergillus foetidus ZU-G1 using response surface methodology. Journal of Food Science 72(4):120-125.
Michelena, V.V. and F.J. Castillo. 1984. Production of amylase by Aspergillus foetidus on rice flour medium and characterization of the enzyme. Journal of Applied Bacteriology 56(3): 395-407.
Mitchell, D.B., K. Vogel, B.J. Weimann, L. Pasamontes and A.P.G.M. van Loon. 1997. The phytase subfamily of histidine acid phosphatase: isolation of genes for two novel phytases from fungi Aspergillus terreus andMyceliophthora thermophila. Microbiology 143(1): 245-252.
Moslem, M., K. Abd-Elsalam, M. Yassin and A. Bahkali. 2010. First morphomolecular identification of Penicillium griseofulvum and Penicillium aurantiogriseum toxicogenic isolates associated with blue mold on apple. Foodborne Pathogens and Disease 7(7):857-861.
Murry, A.C., R.D. Lewis and H.E. Amos. 1997. The effect of microbial phytase in a pearl millet-soybean meal diet on apparent digestibility and retention of nutrients, serum mineral concentration, and bone mineral density of nursery pigs. Journal of Animal Science 75(5): 1284-1291.
Oh, B.C., W.C. Choi, S. Park, Y.O. Kim and T.K. Oh. 2004. Biochemical properties and substrate specificities of alkaline and histidine acid phytases. Applied Microbiology and Biotechnology 63(4): 362-72.
Osorio, N.W. and M. Habte. 2001. Synergistic influence of an arbuscular mycorrhizal fungus and a P solubilizing fungus on growth and P uptake of Leucaena leucocephala in an oxisol. Arid Land Research and Management 15(3): 263-274.
Pasamontes, L., M. Haiker, M. Wyss, M. Tessier and A.P.G.M. van Loon. 1997. Gene cloning, purification, and characterization of a heat-stable phytase from the fungus Aspergillus fumigatus. Applied and Environmental Microbiology 63(5): 1696-1700.
Payne, G.A., W.C. Nierman, J.R. Wortman, B.L. Pritchard, D. Brown, R.A. Dean, D. Bhatnagar, T.E. Cleveland, M. Machida and J. Yu. 2006. Whole genome comparison of Aspergillus flavus and A. oryzae. Medical Mycology 44 (Suppl 1): 9-11.
Peterson, S.W., E.M Bayer and D.T. Wicklow. 2004. Penicillium thiersii, Penicillium angulare and Penicillium decaturense, new species isolated from wood-decay fungi in North America and their phylogenetic placement from multilocus DNA sequence analysis. Mycologia 96(6): 1280-1293.
Purohit, J.S., J.R. Dutta, R.K. Nanda and R. Banerjee. 2006. Strain improvement for tannase production from co-culture of Aspergillus foetidus and Rhizopus oryzae. Bioresource Technology 97(6): 795-801.
Raboy, V. 2003. myo-inositol-1,2,3,4,5,6-hexakisphosphate. Phytochemistry 64(6): 1033-1043.
Seifert, K., G. Morgan-Jones, W. Gams and B. Kendrick. 2011. The genera of hyphomycetes. APS press, MN, USA. 997 p.
Shieh, T.R. and J.H. Ware. 1968. Survey of microorganisms for the production of extracellular phytase. Applied Microbiology 16(9): 1348-1351.
Shieh, T.R., R.J. Wodzinski and J.H. Ware. 1969. Regulation of the formation of acid phosphatase by inorganic phosphate in Aspergillus ficuum. Journal of Bacteriology 100(3): 1161-1165.
Staheli, J.P., R. Boyce, D. Kovarik and T.M. Rose. 2011. CODEHOP PCR and CODEHOP PCR primer design. Methods in Molecular Biology 687: 57-73.
Steinera, T., M. Rainer, F. Ariane and J.S. Jakobb. 2006. Influence of feeding level on apparent total tract digestibility of phosphorus and calcium in pigs fed low-phosphorus diets supplemented with microbial or wheat phytase. Livestock Science 102(1): 1-10.
Tamura, K., J. Dudley, M. Nei and S. Kumar. 2007. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Molecular Biology and Evolution 24: 1596-1599.
Thompson, J.D., D.G. Higgins and T.J. Gibson. 1994. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Research 22(22):4673-4680.
Tseng, Y.H., T.J. Fang and S.M. Tseng. 2000. Isolation and characterization of a novel phytase from Penicillium simplicissimum. Folia Microbiologica 45(2): 121-127.
Tyagunova, Y. and J.L. Sorensen. 2010. The bio-transformation of aromatic amino acids by Phoma macrostoma. Natural Products Communications 5(1): 81-84.
UniProt Consortium. 2008. The universal protein resource (UniProt). Nucleic Acids Research 36 (Database issue): D190-5.
Ullah, A.H. and B.Q. Phillippy. 1994. Substrate selectivity in Aspergillus ficuum phytase and acid phosphatases using myo-inositol phosphates. Journal of Agricultural and Food Chemistry 42(2): 423-425.
Valte, R.D., P.C. Borude, A.K. Hule and A.R. Juvekar. 2010. Optimization of xylanase production from Aspergillus foetidus MTCC 4898 by solid state fermentation using statistical methods. Journal of General and Applied Microbiology 56(1): 75-80.
Vats, P. and U.C. Banerjee. 2004. Production studies and catalytic properties of phytases (myo-inositolhexakisphosphate phosphohydrolases): an overview. Enzyme and Microbial Technology 35(1): 3-14.
Wakiyama, M., H. Tanaka, K. Yoshihara, S. Hayashi and K. Ohta. 2008. Purification and properties of family-10 endo-1,4-beta-xylanase from Penicillium citrinum and structural organization of encoding gene. Journal of Bioscience and Bioengineering 105(4): 367-374.
Wang, Y.N., L. Tian, H.M. Hua, X. Lu, S. Sun, H.H. Wu and Y.H. Pei. 2009. Two new compounds from the broth of the marine fungus Penicillium griseofulvum Y19-07. Journal of Asian Natural Products Research 11(11): 912-917.
Webster, J. and R.W.S. Weber. 2007. Introduction to fungi. Cambridge University Press, UK. 841 p.
White, T.J., T. Bruns, S. Lee and J.W. Taylor. 1990. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. pp. 315-322. In: Innis, M.A., D.H. Gelfand, H.H. Sninsky and T.J. White. (eds.). PCR Protocols: A Guide to Methods and Applications. Academic Press, New York. 482 p.
Wodzinski, R.J. and A.H.J. Ullah. 1996. Phytase. Advances in Applied Microbiology 42: 263-302.
Xiong, A.S., Q.H. Yao, R.H. Peng, X. Li, H.Q. Fan, M.J. Guo and S.L. Zhang. 2004. Isolation, characterization, and molecular cloning of the cDNA encoding a novel phytase from Aspergillus niger 113 and high expression in Pichia pastoris. Journal of Biochemistry and Molecular Biology 37(3): 282-291.
Xiong, A.S., Q.H. Yao, R.H. Peng, P.L. Ha, Z.M. Cheng and Y. Li. 2005. High level expression of a recombinant acid phytase gene in Pichia pastoris. Journal of Applied Microbiology 98: 418-428.
Yadava, B.K. and J.C. Tarafdara. 2011. Penicillium purpurogenum, unique P mobilizers in arid agro-ecosystems. Arid Land Research and Management 25(1): 87-99.
Zhang, Y., C. Li, D.C. Swenson, J.B. Gloer, D.T. Wicklow and P.F. Dowd. 2003. Novel antiinsectan oxalicine alkaloids from two undescribed fungicolous Penicillium spp. Organic Letters 5(5):773-776.
Zhang, G.Q., X.F. Dong, Z.H. Wang, Q. Zhang, H.X. Wang and J.M. Tong. 2010. Purification, characterization, and cloning of a novel phytase with low pH optimum and strong proteolysis resistance from Aspergillus ficuumNTG-23. Bioresource Technology 101(11): 4125-4131.
Zhang, H., X. Wu, L. Gang and Q. Pei. 2011. Interactions between arbuscular mycorrhizal fungi and phosphate-solubilizing fungus (Mortierella sp.) and their effects on Kostelelzkya virginica growth and enzyme activities of rhizosphere and bulk soils at different salinities. Biology and Fertility of Soils 47: 543-554.
Zhao, Q., H. Liu, Y. Zhang and Y. Zhang. 2010. Engineering of protease-resistant phytase from Penicillium sp.: high thermal stability, low optimal temperature and pH. Journal of Bioscience and Bioengineering 110(6): 638-645.
How to Cite
APA
ACM
ACS
ABNT
Chicago
Harvard
IEEE
MLA
Turabian
Vancouver
Download Citation
Article abstract page views
Downloads
License
Copyright (c) 2015 Revista Facultad Nacional de Agronomía

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
The journal allows the author(s) to maintain the exploitation rights (copyright) of their articles without restrictions. The author(s) accept the distribution of their articles on the web and in paper support (25 copies per issue) under open access at local, regional, and international levels. The full paper will be included and disseminated through the Portal of Journals and Institutional Repository of the Universidad Nacional de Colombia, and in all the specialized databases that the journal considers pertinent for its indexation, to provide visibility and positioning to the article. All articles must comply with Colombian and international legislation, related to copyright.
Author Commitments
The author(s) undertake to assign the rights of printing and reprinting of the material published to the journal Revista Facultad Nacional de Agronomía Medellín. Any quotation of the articles published in the journal should be made given the respective credits to the journal and its content. In case content duplication of the journal or its partial or total publication in another language, there must be written permission of the Director.
Content Responsibility
The Faculty of Agricultural Sciences and the journal are not necessarily responsible or in solidarity with the concepts issued in the published articles, whose responsibility will be entirely the author or the authors.