IDENTIFICACIÓN MOLECULAR DE MICROORGANISMOS ASOCIADOS A LA RIZOSFERA DE PLANTAS DE VAINILLA EN COLOMBIA
Molecular Identification of Microorganisms Associated to the Rhizosphere of Vanilla Plants in Colombia
Palabras clave:
ADNr, PCR, PGPR, filogenia, Vanilla planifolia (es)PCR, PGPR, Phylogeny, rDNA, Vanilla planifolia (en)
El cultivo de vainilla es altamente promisorio en Colombia, pero se requiere mayor conocimiento de su manejo agronómico y de los microorganismos que crecen asociados a su rizosfera, de los cuales depende esta planta para su nutrición y crecimiento. En este trabajo se realizaron aislamientos de microorganismos de la rizosfera de plantas de vainilla en un cultivo piloto ubicado en el municipio de Sopetrán (Antioquia, Colombia). Los microorganismos se aislaron en medios selectivos de crecimiento para evaluar su capacidad para descomponer celulosa, proteínas, solubilizar fosfato inorgánico y orgánico (fitato) y fijar nitrógeno en forma asimbiótica. Una vez aislados y purificados, se obtuvieron un total de 109 aislamientos, de los cuales se seleccionaron 52 morfotipos para su identificación molecular por secuenciación de las regiones ITS y 16S del ADN ribosomal para hongos y bacterias, respectivamente. Se encontró una alta variedad de microorganismos en la rizosfera de plantas de vainilla, destacándose las bacterias Bacillus megaterium, Pseudomonas koreensis y Acinetobacter sp. y el hongo Plectosphaerella sp., por su potencial para ser utilizados como biofertilizantes destinados a mejorar la nutrición y el crecimiento de estas plantas.
The cultivation of vanilla (Vanilla planifolia) is highly promising in Colombia, but more research is needed on its agronomical management and beneficial microorganisms that grow associated to its rhizosphere, on which the plant depends for its nutrition and growth. This study involved the identification of microorganisms associated to the rhizosphere of vanilla plants in a crop located in Sopetrán, Colombia. The microbes were isolated in selective media for functional groups such as cellulolytic, proteolytic, inorganic and organic phosphate (phytate) solubilizers, and asymbiotic nitrogen fixing bacteria. After isolation and purification, 109 microbial isolates were obtained. DNA was extracted from 52 selected isolates for molecular identification based on ITS and 16S rDNA sequencing, for fungi and bacteria, respectively. The diversity of rhizosphere microorganisms found was significant. Bacteria such as Bacillus megaterium, Pseudomonas koreensis and Acinetobacter sp., and the fungus Plectosphaerella sp., may have a high potential to be used as biofertilizers to improve vanilla plant nutrition and growth.
IDENTIFICACIÓN MOLECULAR DE MICROORGANISMOS ASOCIADOS A LA RIZOSFERA DE PLANTAS DE VAINILLA EN COLOMBIA
Molecular Identification of Microorganisms Associated to the Rhizosphere of Vanilla Plants in Colombia
CLAUDIA L. ÁLVAREZ1, MSc.; NELSON W. OSORIO1, Ph. D.; MAURICIO MARÍN MONTOYA1, Ph. D.
1 Laboratorios de Biología Celular y Molecular y de Microbiología Industrial, Facultad de Ciencias, Universidad Nacional de Colombia, sede Medellín, Colombia. klaualvarez@gmail.com, nwosorio@unal.edu.co, mamarinm@unal.edu.co. Autor de correspondencia: Prof. Mauricio Marín Montoya. Departamento de Biociencias, Facultad de Ciencias, Universidad Nacional de Colombia, sede Medellín, Colombia. Calle 59 # 63-20, of. 11-113, Medellín, Colombia. A.A. 3840, Medellín. Fax: (+57-4) 430 93 32. mamarinm@unal.edu.co.
Presentado el 19 de febrero de 2013, aceptado el 18 de junio de 2013, correcciones el 28 de junio de 2013.
RESUMEN
El cultivo de vainilla es altamente promisorio en Colombia, pero se requiere mayor conocimiento de su manejo agronómico y de los microorganismos que crecen asociados a su rizosfera, de los cuales depende esta planta para su nutrición y crecimiento. En este trabajo se realizaron aislamientos de microorganismos de la rizosfera de plantas de vainilla en un cultivo piloto ubicado en el municipio de Sopetrán (Antioquia, Colombia). Los microorganismos se aislaron en medios selectivos de crecimiento para evaluar su capacidad para descomponer celulosa, proteínas, solubilizar fosfato inorgánico y orgánico (fitato) y fijar nitrógeno en forma asimbiótica. Una vez aislados y purificados, se obtuvieron un total de 109 aislamientos, de los cuales se seleccionaron 52 morfotipos para su identificación molecular por secuenciación de las regiones ITS y 16S del ADN ribosomal para hongos y bacterias, respectivamente. Se encontró una alta variedad de microorganismos en la rizosfera de plantas de vainilla, destacándose las bacterias Bacillus megaterium, Pseudomonas koreensis y Acinetobacter sp. y el hongo Plectosphaerella sp., por su potencial para ser utilizados como biofertilizantes destinados a mejorar la nutrición y el crecimiento de estas plantas.
Palabras clave: ADNr, Filogenia, PCR, PGPR, Vanilla planifolia.
ABSTRACT
The cultivation of vanilla (Vanilla planifolia) is highly promising in Colombia, but more research is needed on its agronomical management and beneficial microorganisms that grow associated to its rhizosphere, on which the plant depends for its nutrition and growth. This study involved the identification of microorganisms associated to the rhizosphere of vanilla plants in a crop located in Sopetrán, Colombia. The microbes were isolated in selective media for functional groups such as cellulolytic, proteolytic, inorganic and organic phosphate (phytate) solubilizers, and asymbiotic nitrogen fixing bacteria. After isolation and purification, 109 microbial isolates were obtained. DNA was extracted from 52 selected isolates for molecular identification based on ITS and 16S rDNA sequencing, for fungi and bacteria, respectively. The diversity of rhizosphere microorganisms found was significant. Bacteria such as Bacillus megaterium, Pseudomonas koreensis and Acinetobacter sp., and the fungus Plectosphaerella sp., may have a high potential to be used as biofertilizers to improve vanilla plant nutrition and growth.
Keywords: PCR, PGPR, Phylogeny, rDNA, Vanilla planifolia.
INTRODUCCIÓN
La vainilla es la segunda especia de mayor valor en el mundo, después del azafrán, y es el saborizante más utilizado en la industria alimenticia. Este es un cultivo promisorio en diferentes regiones de Colombia que presentan condiciones agroecológicas apropiadas para su crecimiento y desarrollo. Sin embargo, su extensión en el país es muy pequeña (72 ha) y los rendimientos que se obtienen son bajos, aproximadamente un 50 % menos, comparados con otros países como Madagascar, México o Islas Reunión (Damirón, 2004; Ledezma et al., 2006), donde la producción promedio es de 1,2 kg de vainas verdes por planta al año. Esto se debe, entre otros aspectos, al desconocimiento que se tiene en Colombia del manejo agronómico del cultivo, de sus problemas fitosanitarios, necesidades fisiológicas y requerimientos nutricionales (Moreno y Díez, 2011).
La nutrición de la vainilla es uno de los aspectos más críticos para su producción comercial, debido a que está estrechamente ligada con un mejor desarrollo vegetativo de la planta, lo que conlleva a una mayor floración y, por ende, al aumento en la producción de vainas. Como respuesta a una mejor nutrición se han obtenido plantas de vainilla más vigorosas y con mayor floración y fructificación (Castro, 2008). La información al respecto es limitada, pero se sabe que al inicio del cultivo los aportes de materia orgánica vegetal en descomposición (Osorio et al., 2011), y las podas en periodos de seis meses antes de la floración permiten lograr el vigor deseado en las plantas (Damirón, 2004). En nuestro medio se desconocen los niveles óptimos de nutrientes requeridos por la planta de vainilla, por lo que en la práctica son inferidos de especies hortícolas de otros miembros de la familia Orchidaceae (Hernández y Lubinsky, 2010).
En investigaciones recientes en Colombia, Osorio et al. (2011) reportaron que la combinación de 75 % de material orgánico leñoso o fibra de coco, 25 % de hojarasca y 7 g por planta del fertilizante 27-11-11, incrementaron significativamente el crecimiento y desarrollo de plantas de vainilla recién establecidas. En explotaciones comerciales de la India, se realizan aplicaciones anuales por planta de 40 a 60 g de nitrógeno (N), 20 a 30 g de fosforo (P2O5) y 60 a 100 g de potasio (K2O), así como aplicaciones foliares del fertilizante 17-17- 17 una vez por mes, para estimular el crecimiento y floración (Anandaraj et al., 2005). Debido a que el sistema radicular de la vainilla es superficial y sólo se desarrolla directamente sobre la capa de materia orgánica del suelo (Castro, 2008), se considera que los microorganismos rizosféricos juegan un papel importante en la descomposición de los diversos sustratos orgánicos utilizados para su establecimiento. De esta manera, pueden determinar la disponibilidad y toma de nutrientes por las plantas y así promover su crecimiento (Castro, 2008).
Muchos de los microorganismos de la rizosfera de las plantas forman un biocomponente integral de los suelos que controla los ciclos biogeoquímicos de nutrientes. Estos microorganismos se pueden clasificar en grupos funcionales tales como bacterias fijadoras de nitrógeno (FBN), microorganismos solubilizadores de fosfato (PSM), microorganismos celulolíticos (CEL) y amilolíticos, microorganismos proteolíticos (PRO) -amonificantes- y hongos micorrizales, entre otros (Sylvia et al., 1999). En los cultivos de vainilla de Colombia y otros países, es necesario aumentar el conocimiento que se tiene sobre los microorganismos que habitan la rizosfera, así como de sus nichos ecológicos y de los mecanismos de interacción que existen entre los diferentes grupos. Algunos de los reportes disponibles, indican el uso de Pseudomonas fluorescens, Azospirillum y bacterias solubilizadoras de P, como biofertilizantes útiles para promover el crecimiento de vainilla (Anilkumar, 2004). Así mismo, Surendra et al. (2009), desarrollaron un consorcio de PGPR's (Plant Growth Promoting Rhizobacteria) para la micropropagación ex vitro de plantas de vainilla y encontraron que diversas especies de Azotobacter, Pseudomonas y Bacillus fueron las más eficientes.
El objetivo de este estudio fue aislar e identificar microorganismos rizosféricos con potencial para hacer más disponibles los nutrientes en el cultivo de vainilla y así contribuir a un posterior desarrollo de biofertilizantes. Para esto se evaluaron diferentes grupos funcionales y se realizó su identificación molecular a través de la secuenciación de regiones ITS (hongos) y 16S (bacterias) del ADN ribosomal (ADNr).
MATERIALES Y MÉTODOS
AISLAMIENTO DE MICROORGANISMOS
Muestras de la rizosfera de plantas de vainilla de un año de edad se tomaron de diferentes lotes de un cultivo bajo cobertizos de techo-sombra, ubicado en el municipio de Sopetrán (Antioquia, Colombia). El sustrato en el que se establecieron las plantas correspondía a 85 % de hojarasca, 15 % de material leñoso y 5 % de estiércol de cabras. Las muestras de raíces se cortaron con una tijera podadora previamente desinfestada con alcohol al 70 %, se empacaron en bolsas plásticas con el sustrato y se conservaron en refrigeración a 4 °C hasta su análisis posterior.
Para el aislamiento de los microorganismos el material colectado se mezcló homogéneamente sobre papel craft y se tomaron al azar 10 g (base seca). Con esta muestra se prepararon diluciones seriales hasta 10-8 en agua desionizada destilada estéril, en CaCl2 0,01 M y en agua peptonada al 1 %. Posteriormente, se transfirieron 50 µL a cajas de Petri que contenían medios selectivos para aislar microorganismos celulolíticos (CEL) (Wood, 1980), proteolíticos-amonificantes (PROT) (Wood, 1980), fijadores de N2 (FBN) (Döbereiner y Day, 1976) y solubilizadores de fosfato inorgánico (PSM) (Osorio y Habte, 2001) y orgánico (FIT) (Tabatabai, 1982) (Tabla 1). Todos los medios fueron incubados a 28 °C por 72 h en condiciones de luz natural.
Las colonias microbiales que desarrollaron halos en los medios PSM, CEL, PROT y FIT se seleccionaron para ser purificados en cajas de Petri. En el caso de las bacterias se utilizó agar nutritivo (AN) y para los hongos extracto de malta (EM) y papa dextrosa agar (PDA) suplementado con 50 mg mL-1 de penicilina y de sulfato de estreptomicina. Luego, los cultivos se almacenaron a 4 °C hasta su uso posterior. En los diferentes medios, los halos indican que la actividad deseada en un microorganismo se está expresando. Por ejemplo, degradación de celulosa y de proteína (caseína) en los medios CEL y PROT, producción de acidez en el medio PSM y de fosfatasa en el medio FIT. Para el caso de las bacterias con potencial como fijadoras de N2, el desarrollo de colonias en el medio libre de N (NH4+, NO3- y aminoácidos) fue en sí mismo el indicador de la ocurrencia de dicha actividad.
IDENTIFICACIÓN MOLECULAR DE LOS MICROORGANISMOS
Extracción de ácidos nucleicos
Las bacterias obtenidas se sembraron en tubos de ensayo que contenían 5 mL de medio líquido Luria-Bertani (LB) por 24 h en agitación constante, para luego realizar el proceso de extracción de ADN mediante el método del SDS (Sambrook y Rusell, 2001). Los hongos se sembraron en EM líquido y se incubaron en oscuridad a temperatura ambiente (20-24 °C) por 15 días para obtener abundante micelio. Al cabo de este tiempo, se procedió a obtener el micelio mediante filtración con bomba de vacío y se maceró con nitrógeno líquido, para luego extraer su ADN mediante el método del CTAB 3X (Sambrook y Rusell, 2001), o alternativamente utilizando el kit comercial DNeasy plant mini (Qiagen, Alemania). La integridad del ADN extraído fue determinada por electroforesis en gel de agarosa al 1 %, suplementado con 3 µL de bromuro de etidio (10 mg mL-1) en buffer TBE 1X y visualizado en un transiluminador UV con el software Biodoc analyze (Biometra, Alemania). Las concentraciones de ADN obtenidas se calcularon por absorbancia a longitudes de onda de 260 nm y 280 nm, utilizando un Nanodrop 2000C (Thermo, EE. UU.).
Amplificación y secuenciación de regiones ribosomales
La identificación taxonómica de los aislamientos de hongos se realizó con base en la amplificación de las regiones ITS1-5.8S-ITS2 del ADNr, para esto se utilizaron los cebadores ITS1 (5 TCC GTA GGT GAA CCT GCG G 3 ) e ITS4 (5' TCC TCC GCT TAT TGA TAT GC 3') (White et al., 1990). Para el caso de las bacterias se amplificó la subunidad pequeña (16S) del ADNr con los cebadores pA (5' AGA GTT TGA TCC TGG CTC AG 3') y pc5B (5' TAC CTT GTT ACG ACT T 3 ) (Kuske et al., 1997). Las reacciones de PCR se realizaron en un volumentotal de 25 µL conteniendo 0,5 µM de los cebadores, 1 U de Taq ADN polimerasa recombinante (Fermentas, Lituania), 0,2 mM de dNTPs, 1X de buffer de enzima, 1.8 mM de MgCl2, 1 µL de ADN (50 ng µL-1) y agua destilada estéril. En todas las reacciones se incluyó un control negativo. La amplificación se realizó en un termociclador T3 (Biometra) con un programa que consistió de una desnaturalización inicial a 94 °C por 3 min, seguido por 40 ciclos de 94 °C por 30 s, 55 °C (para ITS) y 60 °C (para 16S) por 30 s, 72 °C por 1 min; y una extensión final a 72 °C por 7 min. Luego de la amplificación, se tomaron5 µL de los productos de reacción para separarlos por electroforesis en gel de agarosa al 1,5 %.
Una vez definida la especificidad por el tamaño de los productos amplificados (600 pb), estos se purificaron utilizando los kits QIAquick PCR Purification y QIAquick Gel Extraction (Qiagen), para su secuenciación directa mediante el sistema Big Dye Terminator Cycle Sequencing Ready Reaction (PE Applied Biosystems, EE. UU.) y su corrido en un secuenciador ABI Prism 3730xl de la compañía Macrogen (Corea del Sur). Las secuencias obtenidas con cada cebador, fueron editadas mediante el programa BioEdit 6.0.6 (Hall, 1999), construyéndose secuencias consenso y confirmándose su identidad por comparación con las bases de datos moleculares, mediante BLASTN (http://www.ncbi.nlm.nih.gov/BLAST/Blast.cgi).
Paralelamente, se obtuvieron del GenBank secuencias de las regiones estudiadas de hongos y bacterias relacionadas taxonómicamente con los microorganismos putativamente identificados mediante el BLASTN. Para su alineación con las generadas en esta investigación, se utilizó el programa Clustal W y se realizó un análisis filogenético por el método de Neighbor-Joining, con 1000 iteraciones para determinar los valores de bootstrap. Las distancias genéticas se calcularon por el método de Kimura 2-parámetros, con el programa Mega 5.0 (Tamura et al., 2011).
RESULTADOS
El proceso de aislamiento de los microorganismos en los medios de cultivo selectivos para diferentes grupos funcionales permitió visualizar una alta diversidad microbial asociada a la rizosfera de plantas de vainilla. Los morfotipos coloniales consistieron en 25 aislamientos de potenciales FBN, 6 de FIT, 20 de PSM, 30 de PROT y 28 de CEL, para un total de 109 cepas (9 hongos y 100 bacterias).
IDENTIFICACIÓN MOLECULAR DE LOS MICROORGANISMOS
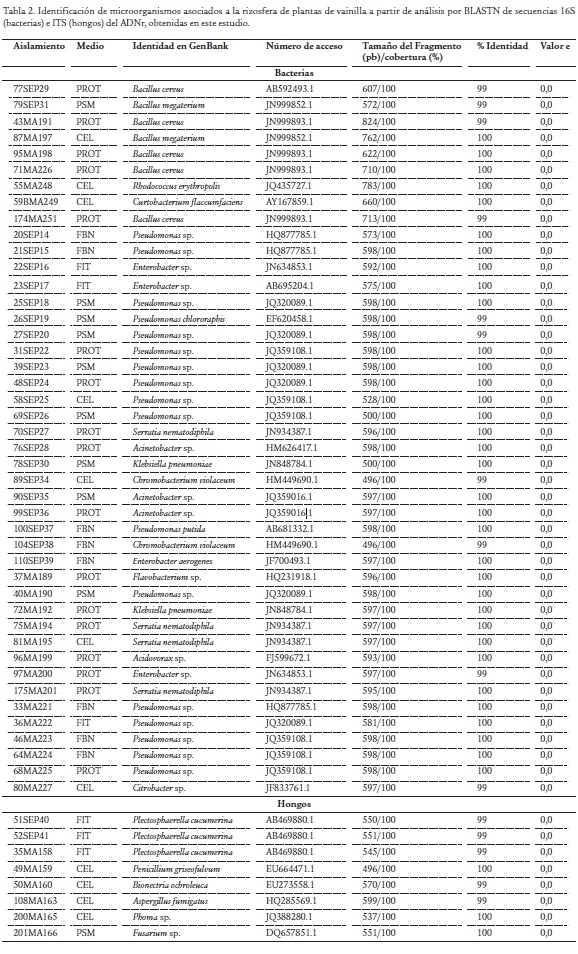
De los 109 aislamientos originalmente obtenidos, se seleccionó para la identificación molecular al menos una cepa de cada morfotipo presente en cada medio de cultivo. De esta manera se evaluaron 44 cepas bacterianas y ocho micóticas (Tabla 2). La amplificación de los productos de PCR de las regiones ITS del ADNr de los hongos generó un fragmento de aproximadamente 600 pb con los cebadores ITS1 e ITS4. Mientras que para la región 16S del ADNr de bacterias, los amplicones obtenidos con los cebadores pA y pc5B tuvieron un tamaño aproximado de 1300-1400 pb.
IDENTIFICACIÓN MOLECULAR DE BACTERIAS
El análisis por BLASTN para las secuencias de bacterias indicó que 17 de las cepas pertenecen al género Pseudomonas, siete a Bacillus, cuatro a Serratia, cuatro a Enterobacter, tres a Acinetobacter, dos a Chromobacterium y dos al género Klebsiella. Además, se halló un representante de los géneros Citrobacter, Acidovorax, Flavobacterium, Rhodococcus y Curtobacterium (Tabla 2). Estos resultados de BLASTN corresponden a un alineamiento local que conduce a identificaciones putativas. Por lo tanto, para tener mayor confiabilidad en la identificación se realizó un análisis filogenético con secuencias de referencia depositadas en GenBank (http://www.ncbi.nlm.nih.gov). Así fue posible en la mayoría de casos la inferencia taxonómica a nivel genérico, aunque para algunas cepas fue posible plantear su hipótesis de identidad a nivel de especie. El análisis filogenético de las bacterias Gram positivas se realizó con base en 742 pb de la región 16S del ADNr e incluyó además de las secuencias obtenidas en este estudio, 23 secuencias representativas de diferentes especies, principalmente de los géneros identificados mediante BLASTN. El dendrograma resultante presentó tres clados principales (I, II y III); el primero de ellos ocupado por representantes del género Bacillus; el segundo clado presentó cepas de referencia de actinomycetes del genéro Streptomyces y el tercero correspondió a corynebacterias de los géneros Curtobacterium, Nocardia y Rhodococcus. De las nueve bacterias Gram positivas obtenidas en el estudio, siete se asociaron filogenéticamente con especies del género Bacillus, siendo dos de ellas (87MA197 y 79SEP31) identificadas como posibles miembros de la especie B. megaterium y las otras cinco (77SEP29, 43MA191, 95MA198, 71MA226, 174MA251), asociadas al complejo de especies B. cereus. Las dos bacterias restantes, se asociaron filogenéticamente con miembros de los géneros Curtobacterium (59BMA249) y Rhodococcus (55MA248) en el clado III, aunque la resolución del análisis no permitió su identificación a nivel de especie (Fig. 1).
La matriz de identidad genética obtenida (no mostrada) arrojó valores de 1 (uno) para las cepas identificadas como B. cereus con representantes de este complejo de especies. Dicho valor fue de 0,997 para las cepas 87MA197 y 79SEP31 con respecto a B. megaterium, lo cual genera un alto soporte para su identificación a nivel de especies. Las primeras bacterias fueron aisladas en su totalidad en el medio que evalúa la capacidad proteolítica, mientras que los dos aislamientos de B. megaterium se obtuvieron de pruebas de celulolíticos y solubilizadores de roca fosfórica (PSM).
Por su parte, el análisis filogenético de las bacterias Gram negativas se realizó con base en 634 pb de la región 16S del ADNr y comprendió las 35 cepas obtenidas en el estudio y 51 secuencias de referencia. El dendrograma resultante presentó cinco clados principales (I a V) soportados por altos valores de bootstrap (>97 %) y diferentes subclados (A, B, C) (Fig. 2). El clado I representa el género Pseudomonas y se subdividió en tres subclados. El IA asoció a 16 de los aislamientos de este trabajo con la especie P. koreensis; el subclado IB presentó cepas de referencia de P. stutzeri, P. fluorescens, P. aeruginosa y P. putida, siendo la cepa 100SEP37 asociada con esta última especie. El subclado IC agrupó a las cepas de referencia de P. syringae.

El segundo clado (II) asoció miembros de los géneros Chromobacterium (subclado IIA), Acidovorax (subclado IIB), y Cupriavidus, Collimonas y Burkholderia (subclado IIIC). En este segundo clado fue posible la identificación de la cepas 89SEP34 y 104SEP38 como miembros del género Chromobacterium y del aislamiento 96MA199 como Acidovorax spp. El clado III correspondió a representantes del género Acinetobacter e incluyó tres de los aislamientos (99SEP36, 90SEP35 y 76SEP28). El clado IV presentó dos subclados (A y B). El grupo IVA permitió identificar a los aislamientos 72MA192 y 78SEP30 como miembros del género Klebsiella, mientras que en el clado IVB se asociaron las cepas 175MA201, 81MA195, 75MA194 y 70SEP27 con Serratia sp.; 97MA200, 23SEP17 y 22SEP16 con el género Enterobacter y la cepa 80MA227 con Citrobacter. Finalmente, en el clado V se ubicaron representantes del género Flavobacterium y la cepa 37MA189 obtenida en este estudio. Los valores de la matriz de identidad genética (no mostrada), soportan la hipótesis taxonómica de que 16 de los aislamientos son afines filogenéticamente a la especie P. koreensis, al compartir un valor de 0,99 con las cepas de referencia. El análisis filogenético y de identidad también permite postular con alto nivel de certeza la naturaleza de las cepas 72MA192 y 75MA194 aisladas en el estudio, porque comparten niveles de identidad de 1 y 0,994 para la región 16S del ADNr con secuencias de referencia de Klebsiella pneumoniae y Enterobacter cloacae, respectivamente.
Por otra parte, los valores de identidad genética obtenidos para las cepas identificadas como Flavobacterium, Acinetobacter, Chromobacterium, así como también de la cepa 100SEP37 asociada con P. putida en el análisis filogenético, indican que estas bacterias solo son identificables en este estudio genérico. De estas bacterias, llaman la atención las dos cepas de Acinetobacter proteolíticas asociadas a la rizosfera de vainilla y un tercer aislamiento de esta especie solubilizador de roca fosfórica.
IDENTIFICACIÓN MOLECULAR DE HONGOS
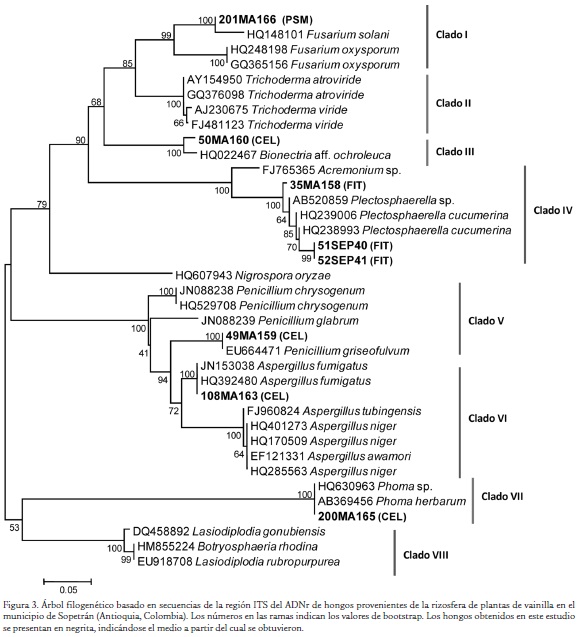
El análisis BLASTN de secuencias ITS indicó que tres de los hongos pertenecen al género Plectosphaerella, mientras que se presentó un representante de los géneros Fusarium, Aspergillus, Penicillium, Bionectria y Phoma (Tabla 2). El análisis filogenético de los hongos obtenidos en la investigación generó un dendrograma con ocho clados (I a VIII) soportados por valores de bootstrap superiores a 94 % (Fig. 3). El primero de ellos representa el género anamórfico Fusarium, y presentó una asociación del aislamiento 201MA166 con la especie fitopatógena F. solani. El clado II correspondió a secuencias de referencia de Trichoderma sp.; mientras que el clado III agrupó a la cepa 50MA160 con un representante del género Bionectria (Hypocreales). El clado IV albergó representantes del ascomycete phyllachoral Plectosphaerella con las cepas 35MA158, 51SEP40 y 52SEP41. Los clados V y VI se presentaron asociados e incluyeron a los géneros anamórficos de hyphomycetes moniliales Penicillium y Aspergillus, siendo las especies más afines filogenéticamente a los aislamientos 49MA159 y 108MA163, Penicillium griseofulvum y Aspergillus fumigatus, respectivamente. El clado VII agrupó al aislamiento 200MA165 con miembros del género anamórfico formador de picnidios (cuerpos fructíferos asexuales) Phoma. Finalmente, el clado VIII correspondió a Botryosphaeria y a algunos de sus estados imperfectos Lasiodiplodia. La matriz de identidad (no mostrada) soportó la asociación genérica de los hongos identificados como Plestosphaerella sp. (0,9), Bionectria sp. (0,92) y Phoma sp. (1). En contraste, el aislamiento 201MA166 asociado en el dendrograma con F. solani y el 49MA159 asociado con P. griseofulvum presentaron bajos niveles de identidad con las secuencias de referencia de dichas especies. Por lo anterior, resulta preferible definir su identidad solo a nivel genérico. Finalmente, el hongo 108MA163 presentó un alto valor (0,99) de identidad con la cepa de referencia de A. fumigatus.
DISCUSIÓN
En este estudio se aislaron e identificaron microorganismos rizosféricos con potencial para hacer más disponibles los nutrientes en el cultivo de vainilla y así contribuir a un posterior desarrollo de biofertilizantes. De las nueve bacterias Gram positivas obtenidas, siete fueron identificadas como pertenecientes al género Bacillus, siendo dos de ellas posibles miembros de la especie B. megaterium y las cinco restantes asociadas al complejo de especies B. cereus. Este complejo, está conformado por las especies B. cereus sensu stricto, B. thuringiensis y B. anthracis, no siendo posible con la resolución que ofrecen las secuencias del ADNr 16S, diferenciar dichas especies. B. anthracis es una bacteria ampliamente reconocida por causar el ántrax y puede ser identificada con base en la amplificación de los genes lef, cya y pag del plásmido pXO1. B. thuringiensis se diferencia de las demás especies, porque tiene la capacidad de producir inclusiones cristalinas de proteína (endotoxinas) durante la esporulación. Jensen et al. (2003) indican que para la identificación molecular de estas especies es necesaria la amplificación de los genes que codifican para las proteínas Cyt y Cry, además de btcap y vip3a, codificantes para proteínas involucradas en las síntesis de la cápsula. De interés resultará en un próximo trabajo la identificación a nivel de especie de las cepas obtenidas en esta investigación, de manera que se evalúe su papel en la rizosfera de las plantas de vainilla.
Por su parte B. megaterium es una bacteria ampliamente investigada que se caracteriza por su capacidad promotora de crecimiento vegetal a partir de la producción de compuestos volátiles, la inhibición del crecimiento de la raíz primaria y el incremento en el número y longitud de pelos radiculares y raíces secundarias (Zou et al., 2010). Esta bacteria ha sido reportada también promoviendo el crecimiento en plántulas de Phaseolus vulgaris, gracias al aumento en la producción de citoquininas en las plantas inoculadas (Ortíz-Castro et al., 2008) y como solubilizadora de fosfatos y biocontroladora de patógenos en Capsicum annuum (Akgûl y Mirik, 2008). En forma similar, López et al. (2010), estudiando la comunidad bacteriana endofítica presente en raíces y semillas de tres variedades de frijol a partir de análisis de secuencias 16S del ADNr, encontraron que B. megaterium se encontraba frecuentemente en las raíces de dos de las variedades evaluadas, y adicionalmente que dichos aislamientos tenían capacidad de solubilizar fitato. Por lo anterior, el aislamiento obtenido de B. megaterium en este estudio presenta un potencial como PGPR en plantas de vainilla y debería ser considerado en el futuro para ser evaluado como biofertilizante bajo condiciones de campo.
El análisis filogenético de las bacterias Gram negativas generó cinco clados principales (I a V), siendo mayoritario el clado I, que representó al género Pseudomonas. Las bacterias de este género están ampliamente distribuidas en los suelos agrícolas y tienen diversas funciones relacionadas con la descomposición de materia orgánica y la promoción del crecimiento de las plantas, aunque algunas pueden tener también efectos patogénicos (Saharan y Nehra, 2011). En este trabajo 16 de los aislamientos fueron identificados como P. koreensis, una especie aislada por primera vez de suelos agrícolas de Corea (Kwon et al., 2003) y taxonómicamente afín al grupo de las Pseudomonas fluorescentes. P. koreensis ha sido reportada como una alternativa promisoria para el biocontrol de Oomycetes que producen zoosporas y que son reconocidos patógenos de diversos cultivos (Hultberg et al., 2010). Kitamura et al. (2010), encontraron que varios representantes del género Pseudomonas, incluyendo a P. koreensis, son capaces de sintetizar una amida del ácido succínico, compuesto que es un promotor del crecimiento de las plantas. Adicionalmente, se ha reportado que miembros del género Pseudomonas aislados de nódulos de raíces de leguminosas, presentan afinidad filogenética con P. putida, P. koreensis y P. fluorescens, entre otros (Issar et al., 2012).
De gran interés resultará evaluar en el futuro el papel principal de esta bacteria en la rizosfera de las plantas de vainilla, ya que fue principalmente detectada creciendo en medio selectivo para bacterias con potencial de fijación de N2 y solubilizadores de P. La capacidad para fijar nitrógeno por parte de bacterias del género Pseudomonas ha sido ampliamente debatida. Sin embargo, recientemente se ha confirmado inequívocamente dicha actividad mediante pruebas bioquímicas en especies como P. stuzeri y P. azotifigens (Desnoues et al., 2003, Hatayama et al., 2005). En adición, Yu et al. (2011), secuenciaron el genoma completo de una cepa de P. stuzeri utilizando el sistema Roche/454 GS FLX, y encontraron la presencia no sólo del gen nifH, sino también de al menos 42 genes que codifican para las proteínas del aparato de desnitrificación, asociado al metabolismo de bacterias asimbióticas fijadoras de nitrógeno. Por esto, en trabajos futuros se podrá confirmar la actividad de fijación biológica de N2 en P. koreensis a partir de pruebas de reducción de acetileno a etileno o mediante la detección directa por PCR del gen nifH.
Por otra parte, el análisis filogenético también permitió asociar con alto nivel de certeza la identidad de dos cepas con secuencias de referencia de Klebsiella pneumoniae y Enterobacter cloacae. Estas bacterias son reconocidas por su condición de saprófitas facultativas, frecuentemente encontradas en aguas, suelos y otros sustratos que tienen contacto con estiércol de animales, pues algunas de sus especies son habitantes comunes del tracto digestivo y urinario de diversos mamíferos. Así, K. pneumoniae, que es clínicamente el más importante representante del género Klebsiella, es un patógeno importante en infecciones nosocomiales que se caracteriza por tener una gran cápsula externa de polisacáridos. Sin embargo, en forma natural está presente en los suelos y puede ser aislada de la superficie de las raíces de algunas plantas. Esta bacteria ha sido muy estudiada como una bacteria diazótrofa de vida libre, pues al menos 30 % de las cepas pueden fijar N2 en condiciones anaeróbicas (Reinhardt et al., 2008). Ya que las cepas de K. pneumoniae sensu stricto asociadas con plantas vivas o en descomposición son en general diferentes de las que están asociadas con infecciones en humanos (las primeras utilizan 5-ketogluconato como única fuente de carbono y carecen de cápsulas) (Brisse et al., 2006), resultará de interés evaluar dichas características en las cepas aquí identificadas, de manera que sea posible definir su viabilidad como potenciales biofertilizantes en cultivos de vainilla.
Con respecto a la otra enterobacteria detectada, E. cloacae, se debe indicar que puede llegar a ser un importante patógeno nosocomial, aunque también se ha encontrado frecuentemente en frutas y vegetales. Junto con las especies E. asburiae y E. aerogenes forman el complejo E. cloacae y con el fin de confirmar inequívocamente el resultado aquí presentado, sería de gran interés secuenciar genes funcionales como RpoB, gyrA y mdh, entre otros. Las infecciones causadas por este patógeno causan la más alta tasa de mortalidad humana comparado con otras infecciones de Enterobacter (Grimont y Grimont, 2006a). Aunque este organismo tiene cepas patógenas de humanos, algunos aislamientos han sido usados como agentes de control biológico de Oomycetes que causan la pudrición de las semillas de pepino tales como Pythium ultimum, y como controlador de algunos insectos plaga (Dijk y Nelson, 2000). Esto representa un posible uso en el manejo de Oomycetes del suelo que afectan la vainilla (ej. Phytophthora meadii).
Dada la utilización de estiércol de caprinos en el sustrato de crecimiento de plantas de vainilla, no resultan sorpresivos dichos aislamientos; por el contrario, estos hallazgos llaman la atención sobre la necesidad de realizar un proceso de compostaje completo con los residuos de origen animal utilizados, de manera que prevalezcan unas condiciones de bioseguridad para los productores y consumidores de vainilla.
Otra bacteria Gram negativa que resultó representativa de los microorganismos proteolíticos obtenidos en el estudio fue Serratia sp. Algunas bacterias del género Serratia se caracterizan por formar el pigmento producido como metabolito secundario prodigiosina, cuya función aún no es bien conocida. Esta pigmentación sólo se presenta en un bajo porcentaje de cultivos aislados y es dependiente de la especie y de las condiciones de incubación. Algunas especies de Serratia se encuentran frecuentemente en aguas, suelos, plantas, material vegetal en descomposición y ocasionalmente en el intestino de diversos animales invertebrados (Grimont y Grimont, 2006b). S. marcescens ha sido ampliamente reportada como una bacteria eficiente en la solubilización de fosfato inorgánico, con potencial para su uso en la extracción de P insoluble en minas naturales de este elemento (Farhat et al., 2009).
Con respecto a los miembros del género Acinetobacter identicados en el estudio, es importante indicar que dichas bacterias se caracterizan por ser saprófitas habitantes de suelos, cuerpos de agua, aguas residuales, alimentos y por no ser fermentativas. Diferentes estudios han indicado que esta bacteria es también una importante PGPR y, por tanto, podría ser formulada como biofertilizante de amplio espectro por su capacidad probada para solubilizar fosfatos, fijar N2 y producir ácido indolacético (AIA) (Indiragandhi et al., 2008; Gulati et al., 2009). Adicionalmente, se ha encontrado que bacterias del género Acinetobacter, además de Pseudomonas, Bacillus, Rhodococcus, Rhizobium y Mycobacterium tienen capacidad de producir AIA (Tsavkelova et al., 2007). El AIA de origen microbiano es altamente relevante en la promoción de la germinación de orquídeas, cuando las cepas bacterianas están en estrecha relación con las semillas (Tsavkelova et al., tigación con las cepas de estos géneros obtenidos en el presente estudio, dados los problemas de latencia que experimentan las semillas de vainilla en nuestro medio.
En relación con las demás bacterias identificadas en el estudio, es de notar la alta diversidad de géneros encontrados, incluyendo Rhodococcus, Curtobacterium, Flavobacterium, Acidovorax y Chromobacterium. Su presencia sugiere la existencia en la rizosfera de vainilla de una compleja red de procesos metabólicos que desempeñan diversas funciones en la descomposición de los sustratos utilizados para el establecimiento y la nutrición de esta planta. Así por ejemplo, se destaca el efecto de Rhodococcus en la producción de sideróforos, que conducen a la formación de complejos solubles de Fe que pueden ser tomados por diversas especies vegetales. De igual manera, se ha confirmado que esta bacteria es solubilizadora de fosfatos mediante el mecanismo de producción de ácidos orgánicos (Saharan y Nehra, 2011).
Con respecto a la identificación de hongos asociados a la rizosfera de plantas de vainilla, se encontraron representantes de los géneros Plectosphaerella, Fusarium, Aspergillus, Penicillium, Bionectria y Phoma, siendo muy llamativo el hecho que algunos de estos (ej. Fusarium y Phoma) pudieran ser posibles fitopatógenos de las raíces de plantas de vainilla. Esto conduce a plantear la necesidad de extremar los cuidados con respecto al origen y tratamiento de los sustratos utilizados para el establecimiento de los cultivos. Un mal uso de estas fuentes puede llevar a la dispersión generalizada de patógenos limitantes del cultivo en el país, lo cual afectaría aún más los planes de su expansión. En este sentido, recientemente Santa et al. (2012) realizaron un estudio tendiente a evaluar los hongos asociados a la pudrición basal de las plantas de vainilla, el principal problema fitosanitario de este cultivo en el mundo (Pinaria et al., 2010), y encontraron que la especie Fusarium oxysporum f. sp. vanillae corresponde al agente causal de dicha enfermedad. Pero adicionalmente, identificaron otros hongos de los géneros Fusarium, Phoma y Lasiodiplodia en plantas con síntomas de necrosis de tallos, lo que coincide plenamente con los resultados de este trabajo. El estudio también identificó a tres de los aislamientos micóticos como miembros del género Plectosphaerella y activos en la solubilización de fitato, lo cual los señala como posibles componentes del proceso de solubilización de las fuentes orgánicas de P presentes en macromoléculas de los sustratos utilizados en este cultivo. El fitato es un derivado del myoinositol presente como fuente de almacenamiento de P en diversos tejidos de plantas y especialmente en semillas (Raboy, 2003). En nuestro conocimiento, este es el primer estudio donde se indica indirectamente la capacidad de este hongo para producir fitasas, lo cual representa un importante hallazgo no sólo por su posible utilización como biofertilizante, sino también por su potencial como fuente de enzimas para la industria de alimentación animal, tal como ocurre con hongos como Aspergillus, Penicillium, Mucor y Rhizopus (Vats y Banerjee, 2004). Adicionalmente a los hongos mencionados, en este trabajo también fue posible la identificación de A. fumigatus, P. griseofulvum y Bionectria sp. Los dos primeros hongos son conocidos saprófitos habitantes de diversos sustratos, donde cumplen importantes funciones como degradadores de materia orgánica y fundamentalmente de los componentes de la pared celular de las plantas, gracias a sus múltiples enzimas hidrolíticas que incluyen celulasas, pectinasas y lacasas (de Vries y Visser, 2001). Bionectria por su parte, es un género de Ascomycetes del orden Hypocreales que incluyen saprófitos degradadores de madera, hojarasca y algunas especies micoparasíticas (Schroers, 2001).
Este estudio representa un primer avance en el reconocimiento de la microflora asociada a la rizosfera de un cultivo de vainilla en Colombia. En él se encontró un alto nivel de diversidad con respecto a los géneros y especies de hongos y bacterias identificados, que brinda una importante fuente de información sobre la utilización potencial de microorganismos como biofertilizantes, controladores biológicos o simplemente aceleradores de la descomposición de sustratos orgánicos. Adicionalmente, se identificaron bacterias con posibles problemas de bioseguridad por su condición dual de patógenas humanas y saprófitas, y posibles hongos fitopatógenos asociados a la rizosfera de plantas de vainilla. Ambas situaciones ameritan realizar esfuerzos para certificarla inocuidad de los sustratos de cultivo y su manejo adecuado en los procesos de compostaje, propios de este tipo de explotaciones agrícolas.
AGRADECIMIENTOS
Esta investigación se financió con recursos del Ministerio de Agricultura y Desarrollo Rural de Colombia (proyecto 082-2008V6151-3701) y de la Universidad Nacional de Colombia sede Medellín, CORANTIOQUIA y BIOANDES Ltda. Se agradece el apoyo administrativo y técnico brindado por María Claudia Díez y Juan Esteban Calle.
BIBLIOGRAFÍA
Akgûl DS, Mirik M. Biocontrol of Phytophthora capsici on pepper plants by Bacillus megaterium strains. J Plant Pathol. 2008;90(1):29-34.
Anandaraj M, Rema J, Sasikumar B, Suseela-Bhai R. Vanilla (Extension pamphlet). Kerala, India: Indian Institute of Spices Research; 2005. p. 11.
Anilkumar AS. Vanilla cultivation: A profitable agri-based enterprise. Kerala call. 2004;1:26-30.
Brisse S, Grimont F, Grimont PA. The genus Klebsiella. En: Dworkin M, Falkow S, Rosenberg E, Schleifer K, Stackebrandt E, editors. The Prokaryotes. A handbook on the Biology of Bacteria. 3rd ed. New York: Springer; 2006. p. 159-196.
CastroBG. Evaluación del cultivo y producción de vainilla en la zona de Papantla, Veracruz, México. [Tesis de Doctorado]: Veracruz: Instituto de Ecología; 2008. p. 93.
Damirón VR. La vainilla y su cultivo. Veracruz: Dirección General de Agricultura y Fitosanitaria del estado de Veracruz;2004. p. 50.
De Vries RP, Visser J. Polysaccharides degradation of plant cell wall enzymes involved in Aspergillus. Microbiol Mol Biol Rev. 2001;65(4):497-522.
Desnoues N, Lin M, Guo X, Ma L, Carreño R, Elmerich C. Nitrogen fixation genetics and regulation in a Pseudomonas stutzeri strain associated with rice. Microbiology. 2003; 149(8):2251-2262.
Dijk KV, Nelson EB. Fatty acid competition as a mechanism by which Enterobacter cloacae suppresses Pythium ultimum sporangium germination and Damping-Off. Appl Environ Microbiol. 2000;66(12):5340-5347. 1976. p. 518-538.
Döbereiner J, Day JM. Associative simbiose in subtropical grasses: characterization of microrganisms and dinitrogens fixing sites. En: Newton WE, Nyman, CJ, editors. Proceedings of the 1st International Symposium on Nitrogen Fixation. Pullman: Washington State University Press; 1976. p. 518-538.
Farhat MB, Farhat A, Bejar W, Kammoun R, Bouchaala K, Fourati A, et al. Characterization of the mineral phosphate solubilizing activity of Serratia marcescens CTM 50650 isolated from the phosphate mine of Gafsa. Arch Microbiol. 2009;191(11):815-824.
Grimont F, Grimont PA. The genus Serratia. En: Dworkin M, Falkow S, Rosenberg E, Schleifer K, Stackebrandt E, editors. The Prokaryotes. A handbook on the Biology of Bacteria. 3rd ed. New York: Springer; 2006a. p. 219-244.
Grimont F, Grimont PA. The genus Enterobacter. En: Dworkin M, Falkow S, Rosenberg E, Schleifer K, Stackebrandt E, editors. The Prokaryotes. A handbook on the Biology of Bacteria. 3rd ed. New York: Springer; 2006b. p. 197-214.
Gulati A, Vyas P, Rahi P, Kasana R. Plant growth-promoting and rhizosphere-competent Acinetobacter rhizosphaerae strain BIHB 723 from the cold deserts of the Himalayas. Curr Microbiol. 2009;58(4):371-377.
Hall TA. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser. 1999;41(1):95-98.
Hatayama K, Kawai S, Shoun H, Ueda Y, NakamurA A. Pseudomonas azotifigens sp. nov., a novel nitrogen-fixing bacterium isolated from a compost pile. Int J Syst Evol Microbiol. 2005;55(4):1539-1544.
Hernández HJ, Lubinsky P. Cultivation Systems. En: OdouxE, Grisoni M, Editors. Vanilla. Boca Raton: CRC Press; 2010. p. 75-95.
Hultberg M, Alsberg T, Khalil S, Alsanius B. Suppression of disease in tomato infected by Pythium ultimum with a biosurfactant produced by Pseudomonas koreensis. Biocontrol. 2010;55(3):435-444.
Indiragandhi P, Anandham R, Madhaiyan M, Sa TM. Characterization of plant growth-promoting traits of bacteria isolated from larval guts of diamondback moth Plutella xylostella (Lepidoptera: Plutellidae). Curr Microbiol. 2008;56(4):327-333.
Issar S, Sharma S, Kumar CD, Kumar GH, Gaur RK. Molecular characterization of Pseudomonas spp. isolated from root nodules of various leguminous plants of Shekhawati Region, Rajasthan, India. Am J Plant Sci. 2012;3(1):60-63.
Jensen Gb, Hansen Bm, Eilenberg J, Mahillon J. The hidden lifestyles of B. cereus and relatives. Environ Microbiol. 2003;5(8):631-640.
Kitamura T, Soejima H, Sugiyama T. U.S. Patent No. 2010/0248314A1. Process for production of succinic acid amide compound. Sumimoto Chemical Company. Tokio: Patent Application Publication; 2010. p. 5.
Kuske CR, Bams SM, Busch JD. Diverse uncultivated bacterial groups from soils of the arid Southwestern United States that are present in many geographic regions. Appl Environ Microbiol. 1997;63(9):3614-3621.
Kwon SW, Kim JS, Park I, Yoon SH, Park DH, Lim C, et al. Pseudomonas koreensis sp. nov., Pseudomonas umsongensis sp. nov. and Pseudomonas jinjuensis sp. nov., novel species from farm soils in Korea. Int J Syst Evol Microbiol. 2003;53(1):21-27.
Ledezma E, Ramirez G, Pino-Benitez N. Forest orchids of theChoco region. Lyonia. 2006;10(1):17-31.
López A, Rogel Ma, Ormeño E, Martínez J, Martínez E. Phaseolus vulgaris seed-borne endophytic community with novel bacterial species such as Rhizobium endophyticum sp. nov. Syst Appl Microbiol. 2010;3(6):322-327.
Moreno F, Díez MC, (Editores). Cultivo de vainilla. Contribuciones para el desarrollo de su cadena productiva en Colombia. Medellín: Universidad Nacional de Colombia;2011. p. 109.
Ortíz R, Valencia E, López J. Plant growth promotion by Bacillus megaterium involves cytokinin signaling. Plant Signal Behav. 2008;3(4):263-265.
Osorio A, Gómez N, Arango D, Moreno F, Díez M, Osorio W. Establecimiento y manejo del cultivo de vainilla. En: Moreno F, Díez MC, editores. Cultivo de vainilla. Contribuciones para el desarrollo de su cadena productiva en Colombia. Medellín: Universidad Nacional de Colombia;2011. p. 45-58.
Osorio W, Habte M. Synergistic influence of an arbuscular mycorrhizal fungus and P solubilizing fungus on growth and plant P uptake of Leucaena leucocephala in an Oxisol. Arid Land Res Mgmt. 2001;15(3):263-274.
Pinaria AG, Liew EC, Burguess LW. Fusarium species associated with vanilla stem rot in Indonesia. Australas Plant Pathol. 2010;39(2):176-183.
Raboy V. Myo-inositol-1,2,3,4,5,6-hexakisphosphate.Phytochemistry. 2003;64(6):1033-1043.
Reinhardt EL, Ramos PL, Manfio GP, Barbosa HR, Pavan C, Moreira-Filho CA. Molecular characterization of nitrogen-fixing bacteria isolated from brazilian agricultural plants at São Paulo state. Braz J Microbiol. 2008;39(3):414-422.
Saharan BS, Nehra V. Plant Growth Promoting Rhizobacteria: A critical review. Life Sci Med Res. 2011(1);21:1-30.
Santa C, Marín M, Díez MC. Identificación del agente causal dela pudrición basal del tallo de vainilla en cultivos bajo cobertizos en Colombia. Rev Mex Micol. 2012;35(1):23-34.
Sambrook J, Russell D. Molecular Cloning: A Laboratory Manual. Cold Spring: Cold Spring Harbor LaboratoryPress; 2001. p. 2231.
Schroers HJ. A monograph of Bionectria (Ascomycota, Hypocreales, Bionectriaceae) and its Clonostachys anamorphs. Stud. Mycol. 2001;46:1-214.
Smith S, Read D. Mycorrhizal simbiosis. 2 ed. Londres: Academic Press; 1997. p. 605.
Surendra GK, Mathew SK, Nazeem PA. Development of plant growth promoting microorganisms consortia technology for ex vitro establishment of micropropagated vanilla (Vanilla planifolia Andr.) and ginger (Zingiber officinale Rosc.). [Citado junio de 2011]; Disponible en: URL: http://www.kauhort.in/Agri%20Microbiology.htm.
Sylvia DM, Fuhrmann JJ, Hartel PG, Zuberer DA. Principles and applications of soil microbiology. New Jersey: Prentice Hall; 1999. p. 550.
Tabatabai MA. Soil enzymes, En: Page AL, Miller RH, Kineey DR, editors. Methods of soil analysis, part two, chemical and microbiological properties. Madison: ASA-SSSA;1982. p. 903-947.
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: Molecular Evolutionary Genetics Analysis using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol Biol Evol.2011;28(10):2439-2442.
Tsavkelova EA, Cherdyntseva TA, Klimova SY, Shestakov AI, Botina SG, Netrusov AI. Orchid-associated bacteria produce indole-3-acetic acid, promote seed germination, and increase their microbial yield in response to exogenous auxin. Arch Microbiol. 2007;188(6):655-664.
Vats P, Banerjee UC. Production studies and catalytic properties of phytases (myo-inositolhexakisphosphate phos-phohydrolases): an overview. Enzyme Microb. Technol.2004;35(1):3-14.
Yu H, Yuan M, Lu W, Yang J, Dai S, Li Q, et al. Complete Genome Sequence of the Nitrogen-Fixing and Rhizosphere- Associated Bacterium Pseudomonas stutzeri Strain DSM4166. J. Bacteriol. 2011;193(3):3422-3423.
White TJ, Bruns T, Lee S, Taylor JW. Amplification and direct sequencing of fungal ribosomal RNA genes for phylo- genetics. En: Innis MA, Gelfand DH, Sninsky JJ, White TJ, editors. PCR Protocols: A guide to methods and appli- cations. New York: Academic Press; 1990. p. 315-322.
Wood PJ. Specify in the interactions of direct dyes with polysaccharides. Carbohydr Res. 1980;85(2):271-287.
Zou C, Li Z, Yu D. Bacillus megaterium strain XTBG34 promotes plant growth by producing 2-pentylfuran. J Microbiol. 2010;48(4):460-466.

Este obra está bajo una licencia de Creative Commons Reconocimiento 3.0 Unported.
Referencias
Akgûl DS, Mirik M. Biocontrol of Phytophthora capsici on pepper plants by Bacillus megaterium strains. J Plant Pathol. 2008;90(1):29-34.
Anandaraj M, Rema J, Sasikumar B, Suseela-Bhai R. Vanilla (Extension pamphlet). Kerala, India: Indian Institute of Spices Research; 2005. p. 11.
Anilkumar AS. Vanilla cultivation: A profitable agri-based enterprise. Kerala call. 2004;1:26-30.
Brisse S, Grimont F, Grimont PA. The genus Klebsiella. En: Dworkin M, Falkow S, Rosenberg E, Schleifer K, Stackebrandt E, editors. The Prokaryotes. A handbook on the Biology of Bacteria. 3rd ed. New York: Springer; 2006. p. 159-196.
CastroBG. Evaluación del cultivo y producción de vainilla en la zona de Papantla, Veracruz, México. [Tesis de Doctorado]: Veracruz: Instituto de Ecología; 2008. p. 93.
Damirón VR. La vainilla y su cultivo. Veracruz: Dirección General de Agricultura y Fitosanitaria del estado de Veracruz;2004. p. 50.
De Vries RP, Visser J. Polysaccharides degradation of plant cell wall enzymes involved in Aspergillus. Microbiol Mol Biol Rev. 2001;65(4):497-522.
Desnoues N, Lin M, Guo X, Ma L, Carreño R, Elmerich C. Nitrogen fixation genetics and regulation in a Pseudomonas stutzeri strain associated with rice. Microbiology. 2003; 149(8):2251-2262.
Dijk KV, Nelson EB. Fatty acid competition as a mechanism by which Enterobacter cloacae suppresses Pythium ultimum sporangium germination and Damping-Off. Appl Environ Microbiol. 2000;66(12):5340-5347. 1976. p. 518-538.
Döbereiner J, Day JM. Associative simbiose in subtropical grasses: characterization of microrganisms and dinitrogens fixing sites. En: Newton WE, Nyman, CJ, editors. Proceedings of the 1st International Symposium on Nitrogen Fixation. Pullman: Washington State University Press; 1976. p. 518-538.
Farhat MB, Farhat A, Bejar W, Kammoun R, Bouchaala K, Fourati A, et al. Characterization of the mineral phosphate solubilizing activity of Serratia marcescens CTM 50650 isolated from the phosphate mine of Gafsa. Arch Microbiol. 2009;191(11):815-824.
Grimont F, Grimont PA. The genus Serratia. En: Dworkin M, Falkow S, Rosenberg E, Schleifer K, Stackebrandt E, editors. The Prokaryotes. A handbook on the Biology of Bacteria. 3rd ed. New York: Springer; 2006a. p. 219-244.
Grimont F, Grimont PA. The genus Enterobacter. En: Dworkin M, Falkow S, Rosenberg E, Schleifer K, Stackebrandt E, editors. The Prokaryotes. A handbook on the Biology of Bacteria. 3rd ed. New York: Springer; 2006b. p. 197-214.
Gulati A, Vyas P, Rahi P, Kasana R. Plant growth-promoting and rhizosphere-competent Acinetobacter rhizosphaerae strain BIHB 723 from the cold deserts of the Himalayas. Curr Microbiol. 2009;58(4):371-377.
Hall TA. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser. 1999;41(1):95-98.
Hatayama K, Kawai S, Shoun H, Ueda Y, NakamurA A. Pseudomonas azotifigens sp. nov., a novel nitrogen-fixing bacterium isolated from a compost pile. Int J Syst Evol Microbiol. 2005;55(4):1539-1544.
Hernández HJ, Lubinsky P. Cultivation Systems. En: OdouxE, Grisoni M, Editors. Vanilla. Boca Raton: CRC Press; 2010. p. 75-95.
Hultberg M, Alsberg T, Khalil S, Alsanius B. Suppression of disease in tomato infected by Pythium ultimum with a biosurfactant produced by Pseudomonas koreensis. Biocontrol. 2010;55(3):435-444.
Indiragandhi P, Anandham R, Madhaiyan M, Sa TM. Characterization of plant growth-promoting traits of bacteria isolated from larval guts of diamondback moth Plutella xylostella (Lepidoptera: Plutellidae). Curr Microbiol. 2008;56(4):327-333.
Issar S, Sharma S, Kumar CD, Kumar GH, Gaur RK. Molecular characterization of Pseudomonas spp. isolated from root nodules of various leguminous plants of Shekhawati Region, Rajasthan, India. Am J Plant Sci. 2012;3(1):60-63.
Jensen Gb, Hansen Bm, Eilenberg J, Mahillon J. The hidden lifestyles of B. cereus and relatives. Environ Microbiol. 2003;5(8):631-640.
Kitamura T, Soejima H, Sugiyama T. U.S. Patent No. 2010/0248314A1. Process for production of succinic acid amide compound. Sumimoto Chemical Company. Tokio: Patent Application Publication; 2010. p. 5.
Kuske CR, Bams SM, Busch JD. Diverse uncultivated bacterial groups from soils of the arid Southwestern United States that are present in many geographic regions. Appl Environ Microbiol. 1997;63(9):3614-3621.
Kwon SW, Kim JS, Park I, Yoon SH, Park DH, Lim C, et al. Pseudomonas koreensis sp. nov., Pseudomonas umsongensis sp. nov. and Pseudomonas jinjuensis sp. nov., novel species from farm soils in Korea. Int J Syst Evol Microbiol. 2003;53(1):21-27.
Ledezma E, Ramirez G, Pino-Benitez N. Forest orchids of theChoco region. Lyonia. 2006;10(1):17-31.
López A, Rogel Ma, Ormeño E, Martínez J, Martínez E. Phaseolus vulgaris seed-borne endophytic community with novel bacterial species such as Rhizobium endophyticum sp. nov. Syst Appl Microbiol. 2010;3(6):322-327.
Moreno F, Díez MC, (Editores). Cultivo de vainilla. Contribuciones para el desarrollo de su cadena productiva en Colombia. Medellín: Universidad Nacional de Colombia;2011. p. 109.
Ortíz R, Valencia E, López J. Plant growth promotion by Bacillus megaterium involves cytokinin signaling. Plant Signal Behav. 2008;3(4):263-265.
Osorio A, Gómez N, Arango D, Moreno F, Díez M, Osorio W. Establecimiento y manejo del cultivo de vainilla. En: Moreno F, Díez MC, editores. Cultivo de vainilla. Contribuciones para el desarrollo de su cadena productiva en Colombia. Medellín: Universidad Nacional de Colombia;2011. p. 45-58.
Osorio W, Habte M. Synergistic influence of an arbuscular mycorrhizal fungus and P solubilizing fungus on growth and plant P uptake of Leucaena leucocephala in an Oxisol. Arid Land Res Mgmt. 2001;15(3):263-274.
Pinaria AG, Liew EC, Burguess LW. Fusarium species associated with vanilla stem rot in Indonesia. Australas Plant Pathol. 2010;39(2):176-183.
Raboy V. Myo-inositol-1,2,3,4,5,6-hexakisphosphate.Phytochemistry. 2003;64(6):1033-1043.
Reinhardt EL, Ramos PL, Manfio GP, Barbosa HR, Pavan C, Moreira-Filho CA. Molecular characterization of nitrogen-fixing bacteria isolated from brazilian agricultural plants at São Paulo state. Braz J Microbiol. 2008;39(3):414-422.
Saharan BS, Nehra V. Plant Growth Promoting Rhizobacteria: A critical review. Life Sci Med Res. 2011(1);21:1-30.
Santa C, Marín M, Díez MC. Identificación del agente causal dela pudrición basal del tallo de vainilla en cultivos bajo cobertizos en Colombia. Rev Mex Micol. 2012;35(1):23-34.
Sambrook J, Russell D. Molecular Cloning: A Laboratory Manual. Cold Spring: Cold Spring Harbor LaboratoryPress; 2001. p. 2231.
Schroers HJ. A monograph of Bionectria (Ascomycota, Hypocreales, Bionectriaceae) and its Clonostachys anamorphs. Stud. Mycol. 2001;46:1-214.
Smith S, Read D. Mycorrhizal simbiosis. 2 ed. Londres: Academic Press; 1997. p. 605.
Surendra GK, Mathew SK, Nazeem PA. Development of plant growth promoting microorganisms consortia technology for ex vitro establishment of micropropagated vanilla (Vanilla planifolia Andr.) and ginger (Zingiber officinale Rosc.). [Citado junio de 2011]; Disponible en: URL: http://www.kauhort.in/Agri%20Microbiology.htm.
Sylvia DM, Fuhrmann JJ, Hartel PG, Zuberer DA. Principles and applications of soil microbiology. New Jersey: Prentice Hall; 1999. p. 550.
Tabatabai MA. Soil enzymes, En: Page AL, Miller RH, Kineey DR, editors. Methods of soil analysis, part two, chemical and microbiological properties. Madison: ASA-SSSA;1982. p. 903-947.
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: Molecular Evolutionary Genetics Analysis using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol Biol Evol.2011;28(10):2439-2442.
Tsavkelova EA, Cherdyntseva TA, Klimova SY, Shestakov AI, Botina SG, Netrusov AI. Orchid-associated bacteria produce indole-3-acetic acid, promote seed germination, and increase their microbial yield in response to exogenous auxin. Arch Microbiol. 2007;188(6):655-664.
Vats P, Banerjee UC. Production studies and catalytic properties of phytases (myo-inositolhexakisphosphate phos-phohydrolases): an overview. Enzyme Microb. Technol.2004;35(1):3-14.
Yu H, Yuan M, Lu W, Yang J, Dai S, Li Q, et al. Complete Genome Sequence of the Nitrogen-Fixing and Rhizosphere- Associated Bacterium Pseudomonas stutzeri Strain DSM4166. J. Bacteriol. 2011;193(3):3422-3423.
White TJ, Bruns T, Lee S, Taylor JW. Amplification and direct sequencing of fungal ribosomal RNA genes for phylo- genetics. En: Innis MA, Gelfand DH, Sninsky JJ, White TJ, editors. PCR Protocols: A guide to methods and appli- cations. New York: Academic Press; 1990. p. 315-322.
Wood PJ. Specify in the interactions of direct dyes with polysaccharides. Carbohydr Res. 1980;85(2):271-287.
Zou C, Li Z, Yu D. Bacillus megaterium strain XTBG34 promotes plant growth by producing 2-pentylfuran. J Microbiol. 2010;48(4):460-466.
Cómo citar
APA
ACM
ACS
ABNT
Chicago
Harvard
IEEE
MLA
Turabian
Vancouver
Descargar cita
Visitas a la página del resumen del artículo
Descargas
Licencia
Derechos de autor 2013 Acta Biológica Colombiana

Esta obra está bajo una licencia internacional Creative Commons Atribución 4.0.
1. La aceptación de manuscritos por parte de la revista implicará, además de su edición electrónica de acceso abierto bajo licencia Attribution-NonCommercial-ShareAlike 4.0 (CC BY NC SA), la inclusión y difusión del texto completo a través del repositorio institucional de la Universidad Nacional de Colombia y en todas aquellas bases de datos especializadas que el editor considere adecuadas para su indización con miras a incrementar la visibilidad de la revista.
2. Acta Biológica Colombiana permite a los autores archivar, descargar y compartir, la versión final publicada, así como las versiones pre-print y post-print incluyendo un encabezado con la referencia bibliográfica del articulo publicado.
3. Los autores/as podrán adoptar otros acuerdos de licencia no exclusiva de distribución de la versión de la obra publicada (p. ej.: depositarla en un archivo telemático institucional o publicarla en un volumen monográfico) siempre que se indique la publicación inicial en esta revista.
4. Se permite y recomienda a los autores/as difundir su obra a través de Internet (p. ej.: en archivos institucionales, en su página web o en redes sociales cientificas como Academia, Researchgate; Mendelay) lo cual puede producir intercambios interesantes y aumentar las citas de la obra publicada. (Véase El efecto del acceso abierto).