Publicado
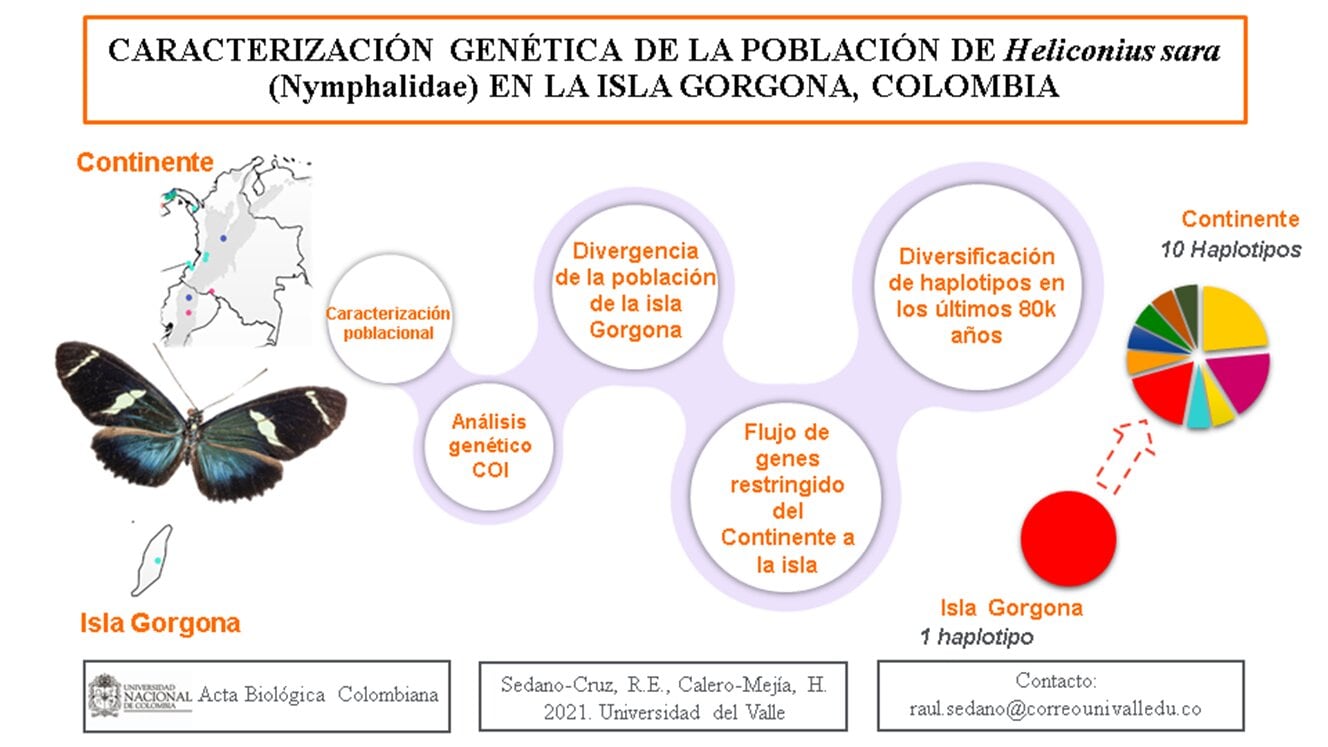
CARACTERIZACIÓN GENÉTICA DE LA POBLACIÓN DE Heliconius sara (Nymphalidae) EN LA ISLA GORGONA, COLOMBIA
Genetic characterization of the Heliconius sara (Nymphalidae) population in the Gorgona Island, Colombia
DOI:
https://doi.org/10.15446/abc.v26n3.86205Palabras clave:
aislamiento por distancia, filogeografía, lepidóptera, COI, mariposas (es)isolation by distance, Phylogeography, lepidoptera, COI, butterflies (en)
Descargas
La estructura genética de poblaciones de mariposas con distribución en islas y sus pares continentales ha sido poco documentada para el neotrópico. Este estudio presenta la caracterización de una población de Heliconius sara con distribución en la Isla Gorgona, ubicada en la región del Pacífico Oriental Colombiano. Para esto se examinaron secuencias parciales de un marcador mitocondrial incluyendo información obtenida del GenBank. Se comparó la diversidad y estructura genética con sus conespecíficos continentales y también con congéneres, con los que comparte un ancestro común cercano en el clado Sapho-Sara. Para el análisis de diversidad y estructura genética se realizó un análisis molecular de varianza. Este análisis muestra que la distancia entre la población de la isla y sus pares en el continente es consistente con la variación intraespecífica observada en otras especies del género Heliconius. Para la reconstrucción de la genealogía y datación reciente en el Pleistoceno superior del grupo monofilético de secuencias de H. sara, se realizó un análisis de inferencia bayesiana, así como una de máxima verosimilitud. Del análisis demográfico se seleccionó un modelo histórico de flujo asimétrico desde la isla hacia el continente que sugiere baja resistencia de la discontinuidad geográfica a la dispersión de esta mariposa diurna desde la isla. Este es el primer estudio en examinar un posible evento de aislamiento de una población insular de mariposas en Colombia.
The genetic structure of butterfly populations among islands and mainland has been poorly documented for the neotropics. This study shows a characterization of the Heliconius sara population with distribution on Gorgona Island in the Colombian Eastern Tropical Pacific region. We obtained partial sequences of a mitochondrial DNA, including information obtained from GenBank. The genetic diversity and structure were compared among the island population and their mainland conspecific, but also with congenerics, with those shared by a recent common ancestor within the Sapho / Sara clade. For the analysis of diversity and genetic structure, an analysis of molecular variance was performed. This analysis shows that the genetic distance between the island's population and that of the mainland is consistent with the intraspecific variation observed in other species of the Heliconious genus. For the reconstruction of the genealogy and the recent dating calibration in the upper Pleistocene of the monophyletic group of H. sara, a Bayesian inference was carried out as well as one of maximum likelihood. From the demographic analysis, an asymmetric gene flow model from the island to the mainland was selected. This model suggests low historical resistance of the geographic discontinuity to dispersal of this small and diurnal butterfly from the island. This is the first study to examine a possible event of local isolation of an island population of a butterfly in Colombia.
Referencias
Arias CF, Salazar C, Rosales C, Kronforst MR, Linares M, Bermingham E, et al. Phylogeography of Heliconius cydno and its closest relatives: disentangling their origin and diversification. Mol Ecol. 2014;23(16):4137-52. Doi: https://doi.org/10.1111/mec.12844
Beerli P, Mashayekhi S, Sadeghi M, Khodaei M, Shaw K. Population Genetic Inference With MIGRATE. Curr Protoc Bioinforma. 2019;68(1):1-28. Doi: https://doi.org/10.1002/cpbi.87
Van Belleghem SM, Baquero M, Papa R, Salazar C, McMillan WO, Counterman BA, et al.. Patterns of Z chromosome divergence among Heliconius species highlight the importance of historical demography. Mol Ecol. 2018;27(19):3852-72. Doi: https://doi.org/10.1111/mec.14560
Beltrán M, Jiggins CD, Brower AVZ, Bermingham E, Mallet J. Do pollen feeding, pupal-mating and larval gregariousness have a single origin in Heliconius butterflies? Inferences from multilocus DNA sequence data. Biol J Linn Soc. 2007;92(2):221-39. Doi: https://doi.org/10.1111/j.1095-8312.2007.00830.x
Beltrán M, Jiggins CD, Bull V, Linares M, Mallet J, McMillan WO, et al. Phylogenetic Discordance at the Species Boundary: Comparative Gene Genealogies Among Rapidly Radiating Heliconius Butterflies. Mol Biol Evol. 2002;19(12):2176-90. Doi: https://doi.org/10.1093/oxfordjournals.molbev.a004042
Benson DA, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW. GenBank. Nucleic Acids Res. 2011;39(Suppl1):D32-7. Doi: https://doi.org/10.1093/nar/gkq1079
Blanco JF, Ramírez A, Scatena F. The streams of Gorgona Natural National Park within the global context : an introduction to the special issue. Actual Biol. 2009;31(91):105-10.
Bohonak AJ, Roderick GK. Dispersal of Invertebrates Among Temporary Ponds: are Genetic Estimates Accurate? Isr J Zool. Taylor & Francis. 2001;47(4):367-86. Doi: https://doi.org/10.1560/HM51-9Y1V-QCRQ-CJP5
Brower AVZ. Rapid morphological radiation and convergence among races of the butterfly Heliconius erato inferred from patterns of mitochondrial DNA evolution. Proc Natl Acad Sci. 1994;91(14):6491-5. Doi: https://doi.org/10.1073/pnas.91.14.6491
Brown KS, Emmel TC, Eliazar PJ, Suomalainen E. Evolutionary patterns in chromosome numbers in neotropical Lepidoptera. Hereditas. 1992;117(2):109-25. Doi: https://doi.org/10.1111/j.1601-5223.1992.tb00165.x
Calero-Mejía H, Armbrecht I, Montoya-Lerma J. Mariposas diurnas y nocturnas (Lepidoptera: Papilionoidae, Saturniidae, Sphingidae) del Parque Nacional Natural Gorgona, Pacífico Colombiano. Rev Biol Trop. 2014;62(Suppl1):317-28. Doi: https://doi.org/10.15517/rbt.v62i0.16351
Chazot N, De-Silva DL, Willmott KR, Freitas AVL, Lamas G, Mallet J, et al. Contrasting patterns of Andean diversification among three diverse clades of Neotropical clearwing butterflies. Ecol Evol. 2018;8(8):3965-82. Doi: https://doi.org/10.1002/ece3.3622
Descimon H. La conservation des Parnassius en France: aspects zoogéographiques, écologiques, démographiques et génétiques. Ciudad, País: Editorial; 1995.
DeVries PJ. The butterflies of Costa Rica and their natural history: Papilionidae, Pieridae, Nymphalidae: Papilionodae, Pieridae, Nymphalidae. Princeton: Princeton University Press; 1987. p.
Drummond AJ, Suchard MA, Xie D, Rambaut A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol. 2012;29(8):1969–1973. Doi: https://doi.org/10.1093/molbev/mss075
Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics. 2010;26(19):2460-1. Doi: https://doi.org/10.1093/bioinformatics/btq461
Ehrlich PR, Raven PH. Butterflies and Plants: A Study in Coevolution. Evolution (N Y). 1964;18(4):586. Doi: https://doi.org/10.2307/2406212
Ersts PJ. Geographic Distance Matrix Generator (version 1.2.3). Am Museum Nat. Hist Cent Biodivers Conserv. 2014.
Excoffier L, Lischer HEL. Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour. 2010;10(3):564-7. Doi: https://doi.org/10.1111/j.1755-0998.2010.02847.x
Fowler HW. Fresh-Water Fishes from Northwestern Colombia. Proc Acad Nat Sci Philadelphia. 1994;96:227-48.
Giraldo A, Diazgranados MC, Gutiérez-Landázuri C. Isla Gorgona, enclave estratégico para los esfuerzos de conservación en el Pacífico Oriental Tropical. Rev Biol Trop. 2014;62, 1–12. Doi: https://doi.org/10.15517/RBT.V62I0.15975
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst Biol. 2010;59(3):307-21. Doi: https://doi.org/10.1093/sysbio/syq010
Halbritter DA, Storer CG, Kawahara AY, Daniels JC. Phylogeography and population genetics of pine butterflies: Sky islands increase genetic divergence. Ecol Evol. 2019;9(23):13389-401. Doi: https://doi.org/10.1002/ece3.5793
Johnson M, Zaretskaya I, Raytselis Y, Merezhuk Y, McGinnis S, Madden TL. NCBI BLAST: a better web interface. Nucleic Acids Res. 2008;36(Suppl 1-2):5-9. Doi: https://doi.org/10.1093/nar/gkn201
Kozak KM, Wahlberg N, Neild AFE, Dasmahapatra KK, Mallet J, Jiggins CD. Multilocus Species Trees Show the Recent Adaptive Radiation of the Mimetic Heliconius Butterflies. Syst Biol. 2015;64(3):505-24. Doi: https://doi.org/10.1093/sysbio/syv007
Kronforst MR, Gilbert LE. The population genetics of mimetic diversity in Heliconius butterflies. Proc R Soc B Biol Sci. 2008;275(1634):493-500. Doi: https://doi.org/10.1098/rspb.2007.1378
Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Battistuzzi FU, editor. Mol Biol Evol. 2018;35(6):1547-9. Doi: https://doi.org/10.1093/molbev/msy096
Leigh JW, Bryant D. POPART: full‐feature software for haplotype network construction. Methods Ecol Evol. 2015;6(9):1110-6. Doi: https://doi.org/10.1111/2041-210X.12410
Linares MC, Soto-Calderón ID, Lees DC, Anthony NM. High mitochondrial diversity in geographically widespread butterflies of Madagascar: A test of the DNA barcoding approach. Mol Phylogenet Evol. Elsevier Inc.; 2009;50(3):485-95. Doi: https://doi.org/10.1016/j.ympev.2008.11.008
Mallet J. Dispersal and gene flow in a butterfly with home range behavior: Heliconius erato (Lepidoptera: Nymphalidae). Oecologia. 1986;68(2):210-7. Doi: https://doi.org/10.1007/BF00384789
Mallet J, Beltrán M, Neukirchen W, Linares M. Natural hybridization in heliconiine butterflies: the species boundary as a continuum. BMC Evol Biol. 2007;7(1):28. Doi:https://doi.org/10.1186/1471-2148-7-28
Maresova J, Habel JC, Neve G, Sielezniew M, Bartonova A, et al., Cross-continental phylogeography of two Holarctic Nymphalid butterflies, Boloria eunomia and Boloria selene. PLoS One. 2019;14(3):e0214483. Doi:https://doi.org/10.1371/journal.pone.0214483
Martin SH, Möst M, Palmer WJ, Salazar C, McMillan WO, Jiggins FM, et al., Natural Selection and Genetic Diversity in the Butterfly Heliconius melpomene. Genetics. 2016;203(1):525-41. Doi: https://doi.org/10.1534/genetics.115.183285
Milá B, Warren BH, Heeb P, Thébaud C. The geographic scale of diversification on islands: genetic and morphological divergence at a very small spatial scale in the Mascarene grey white-eye (Aves: Zosterops borbonicus). BMC Evol Biol. 2010;10(1):158. Doi:https://doi.org/10.1186/1471-2148-10-158
Miller MA, Pfeiffer W, Schwartz T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. 2010 Gatew Comput Environ Work. IEEE; 2010 p. 1-8. Doi:https://doi.org/10.1109/GCE.2010.5676129
Miyakawa M, Hosoi M, Kawakita A, Ito-Harashima S, Yagi T, Ishihara M. Genetic variations and phylogeography of the swallowtail butterfly Papilio machaon on the Japanese Islands. Entomol Sci. 2018;21(3):248-59. Doi:https://doi.org/10.1111/ens.12302
Montejo-Kovacevich G, Smith JE, Meier JI, Bacquet CN, Whiltshire-Romero E, Nadeau NJ, et al., Altitude and life-history shape the evolution of Heliconius wings. Evolution (N Y). 2019;73(12):2436-50. Doi: https://doi.org/10.1111/evo.13865
Morris J, Hanly JJ, Martin SH, Van Belleghem SM, Salazar C, Jiggins CD, et al., Deep Convergence, Shared Ancestry and Evolutionary Novelty in the Genetic Architecture of Heliconius Mimicry. Genetics. 2020;216(3):765-780. Doi: https://doi.org/10.1534/genetics.120.303611
De Moura PA, Quek S-P, Cardoso MZ, Kronforst MR. Comparative population genetics of mimetic Heliconius butterflies in an endangered habitat; Brazil’s Atlantic Forest. BMC Genet. 2011;12(1):9. Doi:https://doi.org/10.1186/1471-2156-12-9
Nadeau NJ, Martin SH, Kozak KM, Salazar C, Dasmahapatra KK, Davey JW, et al., Genome-wide patterns of divergence and gene flow across a butterfly radiation. Mol Ecol. 2013;22(3):814-26. Doi: https://doi.org/10.1111/j.1365-294X.2012.05730.x
Pardo-Diaz C, Salazar C, Baxter SW, Merot C, Figueiredo-Ready W, Joron M, et al., Adaptive Introgression across Species Boundaries in Heliconius Butterflies. R.PLoS Genet. 2012;8(6):e1002752. Doi: https://doi.org/10.1371/journal.pgen.1002752
Parnell AJ, Bradford JE, Curran EV, Washington AL, Adams G, Brien MN, et al. Wing scale ultrastructure underlying convergent and divergent iridescent colours in mimetic Heliconius butterflies. J R Soc Interface. 2018;15(141):20170948. Doi:https://doi.org/10.1098/rsif.2017.0948
QGIS.org. QGIS Sistema de Información Geográfica. Proyecto de Fundación Geoespacial de Código Abierto [Internet]; 2020. Available in: https://qgis.org/es/ Cited: 03 ene 2021.
Rangel-Ch. JO. Islas de Gorgona y Gorgonilla. Colomb Divers Biot I. 1995 p. 145-54.
Rstudio Team. RStudio: Integrated Development for R. Ciudad, Pais: RStudio, Inc. Boston, MA; 2016. p. 77.
Salazar C, Jiggins CD, Taylor JE, Kronforst MR, Linares M. Gene flow and the genealogical history of Heliconius heurippa. BMC Evol Biol. 2008;8(1):132. Doi:https://doi.org/10.1186/1471-2148-8-132
Simon C, Frati F, Beckenbach A, Crespi B, Liu H, Flook P. Evolution, weighting, and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved polymerase chain reaction primers. Ann Entomol Soc Am. 1994;87(6):651-701. Doi: https://doi.org/10.1093/aesa/87.6.651
Templeton A. The meaning of species and speciation: a genetic perspective. En: Otte D, Endler J, editores. Speciat its consequenses. Sunderland: Sinauer Associates; 1989 p. 3-27.
Thurman TJ, Seymoure BM. A bird’s eye view of two mimetic tropical butterflies: Coloration matches predator’s sensitivity. J Zool. 2016;298(3):159-68. Doi:https://doi.org/10.1111/jzo.12305
Todisco V, Gratton P, Cesaroni D, Sbordoni V. Phylogeography of Parnassius apollo: hints on taxonomy and conservation of a vulnerable glacial butterfly invader. Biol J Linn Soc. 2010;101(1):169-83. Doi:https://doi.org/10.1111/j.1095-8312.2010.01476.x
Valente L, Phillimore AB, Melo M, Warren BH, Clegg SM, Havenstein K, et al., A simple dynamic model explains the diversity of island birds worldwide. Nature. Springer US; 2020;579(7797):92-6. Doi:https://doi.org/10.1038/s41586-020-2022-5
Zhang W, Dasmahapatra KK, Mallet J, Moreira GRP, Kronforst MR. Genome-wide introgression among distantly related Heliconius butterfly species. Genome Biol. Genome Biol. 2016;17(1). Doi:https://doi.org/10.1186/s13059-016-0889-0
Zink RM, Barrowclough GF. Mitochondrial DNA under siege in avian phylogeography. Mol Ecol. 2008;17(9):2107-21. Doi:https://doi.org/10.1111/j.1365-294X.2008.03737.x
Zuccon A, Zuccon D. MrEnt: an editor for publication-quality phylogenetic tree illustrations. Mol Ecol Resour. 2014;14(5):1090-4. Doi:https://doi.org/10.1111/1755-0998.12253
Cómo citar
APA
ACM
ACS
ABNT
Chicago
Harvard
IEEE
MLA
Turabian
Vancouver
Descargar cita
Licencia
Derechos de autor 2021 Acta Biológica Colombiana

Esta obra está bajo una licencia internacional Creative Commons Atribución-NoComercial-CompartirIgual 4.0.
1. La aceptación de manuscritos por parte de la revista implicará, además de su edición electrónica de acceso abierto bajo licencia Attribution-NonCommercial-ShareAlike 4.0 (CC BY NC SA), la inclusión y difusión del texto completo a través del repositorio institucional de la Universidad Nacional de Colombia y en todas aquellas bases de datos especializadas que el editor considere adecuadas para su indización con miras a incrementar la visibilidad de la revista.
2. Acta Biológica Colombiana permite a los autores archivar, descargar y compartir, la versión final publicada, así como las versiones pre-print y post-print incluyendo un encabezado con la referencia bibliográfica del articulo publicado.
3. Los autores/as podrán adoptar otros acuerdos de licencia no exclusiva de distribución de la versión de la obra publicada (p. ej.: depositarla en un archivo telemático institucional o publicarla en un volumen monográfico) siempre que se indique la publicación inicial en esta revista.
4. Se permite y recomienda a los autores/as difundir su obra a través de Internet (p. ej.: en archivos institucionales, en su página web o en redes sociales cientificas como Academia, Researchgate; Mendelay) lo cual puede producir intercambios interesantes y aumentar las citas de la obra publicada. (Véase El efecto del acceso abierto).



















