Modelamiento in silico de la liasa organomercurial (MerB) de Pseudomonas fluorescens
In silico modeling of Pseudomonas fluorescens organomercurial lyase (MerB)
Modelagem in silico da liase organomercurial (MerB) de Pseudomonas fluorescens
DOI:
https://doi.org/10.15446/rev.colomb.quim.v51n1.98381Palabras clave:
bioinformática, modelado por homología, organomercurial liasa, Pseudomonas fluorescens (es)Bioinformatics, homology modelling, organomercurial lyase, Pseudomonas fluorescens (en)
bioinformática, modelagem de homologia, organomercurial Liasa, Pseudomonas fluorescens (pt)
El modelamiento in silico ha sido de gran contribución en los procesos proteómicos, desarrollando estructuras de las secuencias proteicas ya existentes, que por motivos de altos costos y las diferentes tecnologías necesarias para el desarrollo de estas metodologías, se encuentran deficientes en el número de modelamientos de proteínas disponibles. Entre aquellas secuencias con carencia de estructura proteica se encuentra la proteína liasa organomercurial (MerB) de Pseudomonas fluorescens, importante en la resistencia al mercurio. En el presente artículo se analizó tanto estructural como funcionalmente la proteína MerB en Pseudomonas fluorescens, utilizando la herramienta de la química estructural “modelamiento por homología” mediante plataformas bioinformáticas, con el fin de obtener un modelo que represente la estructura 3D más precisa y que capturen las mejores variantes estructurales entre todas las posibles conformaciones de las proteínas en la familia. En este trabajo, se desarrolló un método comparativo de la secuencia estudiada con las reportadas en las bases de datos para las proteínas MerB del género Pseudomonas. Se propone un modelo tridimensional para la enzima (MerB) en P. fluorescens, mediante el modelamiento por homología, se muestra la caracterización en la estructura secundaria, terciaria, la caracterización del dominio catalítico y los motivos estructurales presentes.
In silico modeling has made a great contribution to proteomic processes, developing structures of the already existing protein sequences, which for reasons of high costs and the different technologies necessary for the development of these methodologies, are deficient in the number of models of available proteins. Among those sequences lacking protein structure is the organomercurial lyase (MerB) protein from Pseudomonas fluorescens, important in mercury resistance. In this article, the MerB protein in Pseudomonas fluorescens was analyzed both structurally and functionally, using the structural chemistry tool “homology modeling” using bioinformatic platforms, in order to obtain a model that represents the most accurate 3D structure and that captures the best structural variants among all the possible conformations of the proteins in the family. In this work, a comparative method of the sequence studied with those reported in the databases for MerB proteins of the genus Pseudomonas was developed. A three-dimensional model for the enzyme (MerB) in P. fluorescens is proposed, through homology modeling, the characterization at the secondary and tertiary structure level, the characterization of the catalytic domain and the structural motifs present is shown.
A modelagem in silico tem dado um grande contributo para os processos proteómicos, desenvolvendo estruturas de sequências de proteínas já existentes, as quais, pelos elevados custos e pelas diferentes tecnologias necessárias ao desenvolvimento destas metodologias, são deficientes no número de modelos de proteínas disponíveis. Entre as sequências sem estrutura protéica está a proteína organomercurial liase (MerB) de Pseudomonas fluorescens, importante na resistência ao mercúrio. Neste artigo, a proteína MerB em Pseudomonas fluorescens foi analisada estrutural e funcionalmente, usando a ferramenta de química estrutural “modelagem de homologia” usando plataformas de bioinformática, a fim de obter um modelo que represente a estrutura 3D mais precisa e que capture as melhores variantes estruturais. entre todas as conformações possíveis das proteínas da família. Neste trabalho, foi desenvolvido um método comparativo da sequência estudada com aqueles relatados em bancos de dados para proteínas MerB do gênero Pseudomonas. Um modelo tridimensional para a enzima (MerB) em P. fluorescens é proposto, através de modelagem por homologia, a caracterização em nível de estrutura secundária e terciária, a caracterização do domínio catalítico e os motivos estruturais presentes são mostradas.
Orgánica y Bioquímica
Modelamiento in silico de la liasa organomercurial (MerB) de Pseudomonas fluorescens
In silico modeling of Pseudomonas fluorescens organomercurial lyase (MerB)
Modelamiento in silico de la liasa organomercurial (MerB) de Pseudomonas fluorescens
Revista Colombiana de Química, vol. 51, núm. 1, pp. 14-23, 2022
Universidad Nacional de Colombia
Recepción: 12 Septiembre 2021
Revisado: 12 Marzo 2022
Aprobación: 03 Mayo 2022
Resumen: El modelamiento in silico ha sido de gran contribución en los procesos proteómicos, desarrollando estructuras de las secuencias proteicas ya existentes, que por motivos de altos costos y las diferentes tecnologías necesarias para el desarrollo de estas metodologías, se encuentran deficientes en el número de modelamientos de proteínas disponibles. Entre aquellas secuencias con carencia de estructura proteica se encuentra la proteína liasa organomercurial (MerB) de Pseudomonas fluorescens, importante en la resistencia al mercurio. En el presente artículo se analizó tanto estructural como funcionalmente la proteína MerB en Pseudomonas fluorescens, utilizando la herramienta de la química estructural “modelamiento por homología” mediante plataformas bioinformáticas, con el fin de obtener un modelo que represente la estructura 3D más precisa y que capturen las mejores variantes estructurales entre todas las posibles conformaciones de las proteínas en la familia. En este trabajo, se desarrolló un método comparativo de la secuencia estudiada con las reportadas en las bases de datos para las proteínas MerB del género Pseudomonas. Se propone un modelo tridimensional para la enzima (MerB) en P. fluorescens, mediante el modelamiento por homología, se muestra la caracterización en la estructura secundaria, terciaria, la caracterización del dominio catalítico y los motivos estructurales presentes.
Palabras clave: bioinformática, modelado por homología, organomercurial liasa, Pseudomonas fluorescens.
Abstract: In silico modeling has made a great contribution to proteomic processes, developing structures of the already existing protein sequences, which for reasons of high costs and the different technologies necessary for the development of these methodologies, are deficient in the number of models of available proteins. Among those sequences lacking protein structure is the organomercurial lyase (MerB) protein from Pseudomonas fluorescens, important in mercury resistance. In this article, the MerB protein in Pseudomonas fluorescens was analyzed both structurally and functionally, using the structural chemistry tool “homology modeling” using bioinformatic platforms, in order to obtain a model that represents the most accurate 3D structure and that captures the best structural variants among all the possible conformations of the proteins in the family. In this work, a comparative method of the sequence studied with those reported in the databases for MerB proteins of the genus Pseudomonas was developed. A three-dimensional model for the enzyme (MerB) in P. fluorescens is proposed, through homology modeling, the characterization at the secondary and tertiary structure level, the characterization of the catalytic domain and the structural motifs present is shown.
Keywords: Bioinformatics, homology modelling, organomercurial lyase, Pseudomonas fluorescens.
Resumo: A modelagem in silico tem dado um grande contributo para os processos proteómicos, desenvolvendo estruturas de sequências de proteínas já existentes, as quais, pelos elevados custos e pelas diferentes tecnologias necessárias ao desenvolvimento destas metodologias, são deficientes no número de modelos de proteínas disponíveis. Entre as sequências sem estrutura protéica está a proteína organomercurial liase (MerB) de Pseudomonas fluorescens, importante na resistência ao mercúrio. Neste artigo, a proteína MerB em Pseudomonas fluorescens foi analisada estrutural e funcionalmente, usando a ferramenta de química estrutural “modelagem de homologia” usando plataformas de bioinformática, a fim de obter um modelo que represente a estrutura 3D mais precisa e que capture as melhores variantes estruturais. entre todas as conformações possíveis das proteínas da família. Neste trabalho, foi desenvolvido um método comparativo da sequência estudada com aqueles relatados em bancos de dados para proteínas MerB do gênero Pseudomonas. Um modelo tridimensional para a enzima (MerB) em P. fluorescens é proposto, através de modelagem por homologia, a caracterização em nível de estrutura secundária e terciária, a caracterização do domínio catalítico e os motivos estruturais presentes são mostradas.
Palavras-chave: bioinformática, modelagem de homologia, organomercurial Liasa, Pseudomonas fluorescens.
Introducción
Desde hace 20.000-30.000 años A. de C., el mercurio ha sido un elemento incluido en la mano de obra del hombre [1], siendo la minería la más reconocida (exactamente en la extracción de oro a base de mercurio) y, a la vez, la actividad que más contaminación provoca por este metal. Por tanto, durante todo este tiempo ha surgido bioacumulación en su mayor reservorio (ríos, lagos y océanos), provocando afectaciones ambientales y en el ser humano [2]. La bioacumulación en los océanos es debida, por lo general, a las bacterias acuáticas anaeróbicas reductoras de sulfato, que transforman el mercurio elemental a metilmercurio por medio de la biometilación de este elemento [3], siendo el estado más contaminante del mercurio; aunque también puede ser transformado por actividades antropogénicas y ser, posteriormente, llevadas a las zonas acuáticas naturales. Afectando de manera directa a organismos acuáticos, y luego a las personas quienes basan su dieta principalmente de productos marinos [4].
En sintonía con la evolución del bienestar ambiental y humano, han surgido diferentes soluciones para este efecto adverso, como lo es la biorremediación, un mecanismo liderado por microorganismos y sus enzimas, para la degradación y transformación de contaminantes a otro estado de oxidación, el cual es menos tóxico para el medioambiente [5]. Entre estos microorganismos se encuentran las bacterias, uno de cuyos géneros es el Pseudomona [6]; las bacterias de este género son organismos Gram negativos de vida libre, con genomas grandes (en promedio 6 Mb), en los cuales tienen la información para adaptarse fácilmente a las condiciones que el ambiente en el que se encuentra le presente. Una de las especies de este género, con una amplia capacidad de adaptación y que ha sido usada comúnmente en procesos de biorremediación, es el Pseudomonas fluorescens. Además, se ha reportado que este microorganismo ha logrado sobrevivir en lugares con alta concentración de mercurio, por lo que la identificación de los elementos genéticos presentes, y que le dan esta característica, se han convertido en un importante tema de estudio [7].
Se ha identificado que los sistemas de resistencia bacteriana a mercurio se agrupan en un operón, conocido como el “operón Mer”, que varía en estructura y consta de genes que codifican proteínas para la regulación (merR, merD), transporte (merT, merP and/or merC, merF) y reducción de mercurio (MerA) [8] y, en algunos casos, un gen adicional, merB, el cual otorga a los microorganismos que lo presentan la capacidad de cortar el enlace carbono-mercurio de organomercuriales como el acetato fenilmercúrico, y generar Hg2+, que luego es desintoxicado como mercurio metálico por la reductasa mercúrica. Se ha identificado la presencia de este gen en la bacteria P. fluorescens, como uno de los elementos importantes de su capacidad de biorremediación de mercurio [9-11]. MerB es la encargada de escindir el enlace carbono-mercurio por medio de una protonólisis produciendo un compuesto alifático o aromático y Hg (II) [12], el cual posteriormente es reducido a Hg (0) por la reductasa específica MerA [13]. MerB muestra una amplia especificidad de sustrato y puede dividir una gran variedad de compuestos organomercuriales que van desde compuestos de alquilo simples como Me-Hg a poliaromáticos y compuestos heterocíclicos como la merbromina [14]. En diversos reportes se evidencia que, para llevar a cabo esta reacción catalítica, los sitios activos de la proteína MerB varían en los diferentes macroorganismos en los que esta ha sido estudiada, como lo es en el dominio bacteria y en las arqueas. Se han identificado mutaciones génicas que en algunos casos producen cambios en la ubicación de algunos aminoácidos; aunque, en general, en la molécula se mantiene una gran similitud. Sin embargo, se ha identificado que estas variaciones ocasionan cambios en la expresión génica y en la estructura proteica [15].
Son estos últimos elementos los que sustentan la importancia de la biología estructural y la química computacional, con el fin de establecer con exactitud la estructura de las proteínas, confiriendo mayor conocimiento de su estructura, funcionalidad, su dinámica, las interacciones con ligandos y otras proteínas [16, 17]. Tal estudio se realiza por medio de herramientas proteómicas, con el propósito de observar las estructuras proteicas como lo es la cristalización con ayuda de la difracción de rayos X o de resonancia magnética, donde la calidad de la imagen estructural final es directamente proporcional a la perfección de las propiedades físicas del espécimen cristalino, que otorga con precisión las coordenadas atómicas, necesitándose cristales de gran tamaño y calidad suficiente para lograr obtener una recopilación de datos precisa. Sin embargo, este tipo de tecnologías son complejas y muy costosas, por lo que para todas las proteínas identificadas no existen datos de estructura generados experimentalmente [18, 19]. Pero más allá de ser un limitante, también impulsó el desarrollo de la química computacional, la cual, hoy en día, se orienta al modelaje y evaluación de proteínas in silico, una variante viable para ahorrar tiempo en el diseño de experimentos con mejores expectativas de éxito y disminuir costos en materiales y reactivos. Entre estas herramientas se encuentra el “modelado por homología”, también conocido como “modelado comparativo” o “modelado basado en plantilla (TBM)”; estas estrategias de modelado de la estructura 3D de una proteína, usan una estructura conocida experimentalmente como molde.
En el presente artículo se plantea estudiar, con el uso de herramientas bioinformáticas, el gen, la secuencia de proteína y dar a conocer una estrategia para modelar y analizar la estructura tridimensional de la enzima MerB en Pseudomonas fluorescens, la cual no ha sido reportada. De igual manera, se plantea llevar a cabo la caracterización de la estructura secundaria, identificación de dominios funcionales, estructurales y la especificación del centro activo de esta importante molécula básica en los procesos de biorremediación de mercurio.
Materiales y métodos
Secuencias de MerB y alineamiento múltiple
Las secuencias de la familia de MerB en varias especies del género Pseudomonas, se obtuvieron de la plataforma de UniProt [20], estas se usaron para la realización de un alineamiento múltiple con la aplicación del European Bioinformatics Institute [21], T-Coffe [22], con el fin de identificar el nivel de diversidad entre ellas y, por tanto, la homología presente.
Modelamiento de la estructura tridimensional de MerB de Pseudomonas fluorescens
Con la secuencia identificada para MerB de Pseudomona fluorescens, se desarrolló un modelamiento por homología con la aplicación Geno3D [23], donde, en primer lugar, se introduce la secuencia y se corre con los valores exactos que propone la aplicación, con el fin de seleccionar el modelo con mejor valor propuesto, donde luego se extrae su respectiva secuencia y para facilitar su visualización se introduce en el software RasMol [24].
Análisis de la estructura secundaria de MerB de Pseudomonas fluorescens
Se predijo la estructura secundaria de MerB mediante la aplicación PSIPRED [25], la cual utiliza un método altamente preciso para la predicción de la estructura secundaria a partir de secuencias de aminoácidos.
Evaluación y validación de la estructura tridimensional de MerB de Pseudomonas fluorescens
Se evaluó la validación de los modelos predichos usando los programas Biovia [26] para verificar la validez de su estructura secundaria; ProSA-Web [27] para su ajuste a estructuras predichas mediante métodos de laboratorio, como la resonancia magnética nuclear o la difracción de rayos X; y Verify 3D [28], con el fin de llevar a cabo la validación de los modelos de los perfiles tridimensionales.
Identificación de dominios proteicos
Los dominios proteicos de la secuencia de MerB en P. fluorescens, se identificaron por medio de la página web de búsqueda MotifSearch [29].
Resultados
Alineamiento por pares y múltiple
A partir de las secuencias de la familia de las proteínas MerB (Tabla 1), se realizó, en la aplicación Phylogeny.fr [30], el alineamiento múltiple y la obtención del respectivo árbol filogenético, identificando en él las secuencias de MerB para P. fluorescens, y se obtuvieron cuatro secuencias en diferentes posiciones de la filogenia, como se muestra en la Figura 1.
| Código | Organismo |
| Q8G9P0 | Pseudomonas putida 1 (P_putida1) |
| A0A1W6QXX0 | Pseudomonas putida 2 (P_putida2) |
| Q9F3W5 | Pseudomonas sp (P_sp1) |
| O07303 | Pseudomonas sp (P_sp2) |
| Q9Z3Y8 | Pseudomonas sp (P_sp3) |
| Q9F3W6 | Pseudomonas sp (P_sp4) |
| U2A0X4 | Pseudomonas sp (P_sp5) |
| A0A2N7XWW6 | Pseudomonas sp (P_sp6) |
| Q0E6C2| | Pseudomonas aeruginosa (P_aeruginosa1) |
| A0A2N7XXX7| | Pseudomonas sp (P_sp7) |
| A0A212BER9| | Pseudomonas sp (P_sp8) |
| A0A3M5Z239| | Pseudomonas coronafaciens (P_coronafaciens1) |
| A0A1G8QSD9| | Pseudomonas abietaniphila (P_abietaniphila1) |
| A0A5C5R0G9| | Pseudomonas mandelii (P_mandelii1) |
| A0A5P9WB02| | Pseudomonas aeruginosa (P_aeruginosa2) |
| A0A1G5PHN0| | Pseudomonas psychrotolerans (P_psychrotolerans1) |
| A0A2S7FMI2| | Pseudomonas oleovorans (P_oleovorans1) |
| A0A2N7XWV5 | Pseudomonas sp P_sp9 |
| A0A1V0M5B3| | Pseudomonas aeruginosa (P_aeruginosa3) |
| A0A5B0BVV8| | Pseudomonas sp (P_sp10) |
| A0A2N7XX29| | Pseudomonas sp (P_sp11) |
| C0KJV0| | Pseudomonas aeruginosa (P_aeruginosa4) |
| E1V302| | Pseudomonas plecoglossicida (P_plecoglossicida1) |
| A4V7U8| | Pseudomonas fluorescens (P_fluorescens1) |
| S2FX69| | Pseudomonas sp (P_sp12) |
| A0A5C5N7Q8| | Pseudomonas rhodesiae (P_rhodesiae1) |
| A0A5E7MFD1| | Pseudomonas fluorescens (P_fluorescens2) |
| A0A5M8ECB2| | Pseudomonas veronii (P_veronii1) |
| A0A0R3BTV1| | Pseudomonas lactis (P_lactis1) |
| A0A419UJ99| | Pseudomonas synxantha (P_synxantha1) |
| A0A2G0W2S4| | Pseudomonas sp (P_sp13) |
| A0A127HQV2| | Pseudomonas azotoformans (P_azotoformans) |
| A0A0R3B0W4| | Pseudomonas lactis (P_lactis2) |
| A0A3M5ZUV3| | Pseudomonas coronafaciens (P_coronafaciens2) |
| A0A5E7A816| | Pseudomonas fluorescens (P_fluorescens3) |
| Q70MS1| | Pseudomonas fluorescens (P_fluorescens4) |
| E1V304| | Pseudomonas sp (P_sp14) |
| A0A1E4X3J7| | Pseudomonas sp (P_sp15) |
| A0A1T1HZC6| | Pseudomonas sp (P_sp16) |
| E1V303| | Pseudomonas putida (P_putida3) |
| A0A6B7Q571| | Pseudomonas putida (P_putida4) |
| A0A1H0ISK0 | Pseudomonas poael (P_poae1) |
| A0A0P9GCM1 | Pseudomonas syringae (P_syringae1) |
| A0A3S4N2X6| | Pseudomonas putrefaciens (S_putrefaciens2) |
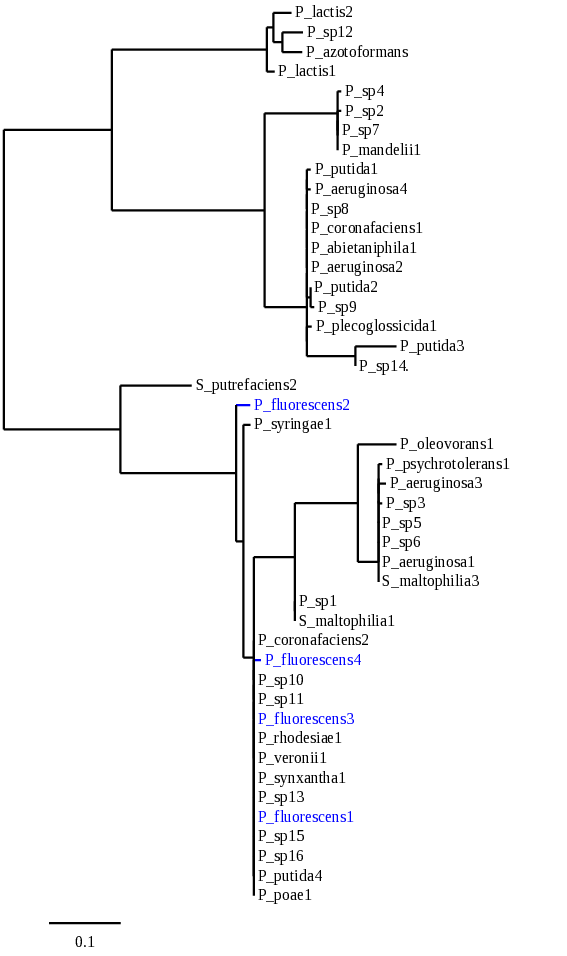
Figura 1. Árbol filogenético producto del alineamiento en Phylogeny.fr [30] de las secuencias reportadas de Pseudomonas para MerB (se resaltan en color azul los aislados de P. fluorescens, en rojo se observan los valores de Bootstrap, con 500 repeticiones, verificando la estabilidad de cada agrupamiento).
A partir del árbol filogenético se observan las diferentes secuencias de MerB de Pseudomona fluorescens, y se identifica la diversidad entre ellas, especialmente la secuencia “P. fluorescens 2”, que tiene mayor similitud con P. syringae, y es diferente a los demás representantes de su especie en, al menos, el 5%. Teniendo en cuenta este nivel de diferencia con las otras secuencias correspondientes en la especie, se escogió la secuencia representativa para MerB de P. fluorescens para el análisis del modelamiento por homología.
Caracterización fisicoquímica de la enzima MerB de P.fluorescens
Partiendo de la secuencia proteica seleccionada de A0A5E7MFD1 para MerB identificada en P. fluorescens(212aa), se verificó por medio de la plataforma Expasy [31] que su peso molecular es de 23139,4 Da y su punto isoeléctrico es de 5,86. Mediante la plataforma Uniprot [20] se estableció su código EC 4.99.1.2, correspondiente a una liasa de alquilmercurio, para la cual su reacción enzimática se presenta en la Figura 2.

Figura 2. Reacción enzimática de MerB identificada para P. fluorescens,dando lugar a una reacción catabólica teniendo como sustratos un alquilmercurio y un protón de hidrógeno, que produce un alcano y un ion mercurio. Tomada de Uniprot [20].
Modelamiento de la estructura tridimensional de la proteína MerB de P. fluorescens
Con los resultados suministrados por Geno3D, se estableció que la mejor secuencia para el modelamiento por homología es la proteína MerB en complejo con mercurio de Escherichia coli 3FN8 (Figura 3). Esta secuencia presentó mayor similitud a la secuencia MerB de P. fluorescens 2 con un valor E de 2 × 10-50 y una desviación media de 0,48 Å.

Figura 3. Modelo 3FN8 computacional generado a partir del estudio por homología para MerB en P. fluorescens a través de RasMol [24], observándose una proteína dimérica, cada cadena con un centro activo de mercurio (esferas rosa) y glicerol (estructura atómica), unidos covalentemente a residuos de aminoácidos.
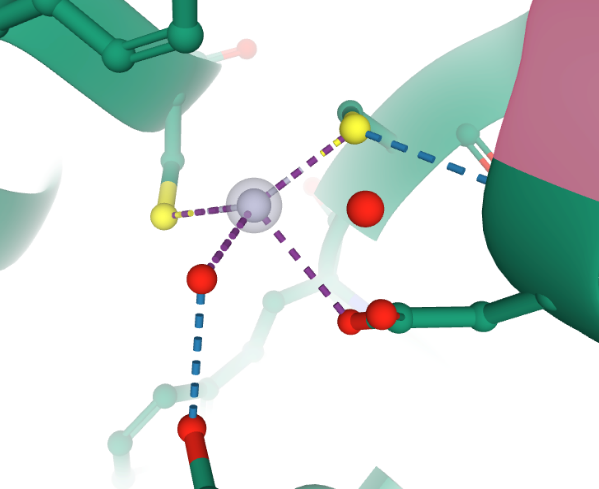
Cada centro activo se logra (observar en la Figura 4) formado por un complejo de mercurio, el cual es clasificado como un complejo metálico, donde se sitúa un ion de mercurio (Hg+2) unido covalentemente mediante enlace metálico a dos cisteínas (Cys 96 y Cys 159) y a un ácido aspártico (Asp 99) y por medio de un puente de hidrógeno a un glicerol (Gol 222).
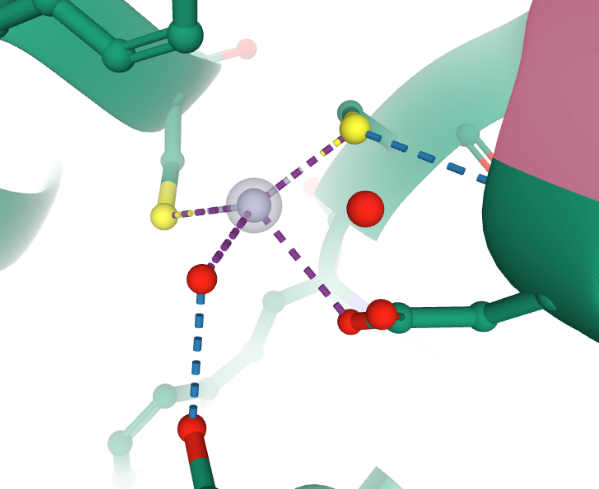
Figura 4. Ilustración computacional del complejo metálico de mercurio para MerB de E. coli 3FN8. Tomada de RCSB PDB Protein Data Bank [32].
Tomando a 3FN8 como molde para el diseño de MerB de P. fluorescens, se generaron cinco modelos (Tabla 2), donde los valores de correlación entre los cinco modelos evaluados se determinaron mediante desviación de la raíz cuadrada media (DRCM) con valores en ángstroms (Tabla 3).
| Nombre del modelo | Molde | Energía del modelo Kcal/mol | Porcentaje de residuos en la región núcleo del diagrama de Ramachandran | Porcentaje de residuos no permitidos en el diagrama de Ramachandran |
| Modelo 1 | 3FN8 | -7.860,73 | 70,3 | 3,8 |
| Modelo 2 | 3FN8 | 4.400,65 | 76,8 | 2,2 |
| Modelo 3 | 3FN8 | 3.824,18 | 73,0 | 1,1 |
| Modelo 4 | 3FN8 | -7.859,05 | 72,4 | 1,6 |
| Modelo 5 | 3FN8 | -7.970,61 | 73,5 | 1,1 |
A partir de los valores obtenidos, se escogió el mejor modelo, el cual corresponde al valor negativo más bajo: el modelo 4; además, tiene porcentajes permitidos en el gráfico de Ramachandran.
| Modelo 1 | Modelo 2 | Modelo 3 | Modelo 4 | Modelo 5 | |
| Modelo 1 | 0,00 | 0,77 | 0,88 | 0,81 | 0,88 |
| Modelo 2 | 0,77 | 0,00 | 0,95 | 0,92 | 0,98 |
| Modelo 3 | 0,88 | 0,95 | 0,00 | 0,88 | 0,87 |
| Modelo 4 | 0,81 | 0,92 | 0,88 | 0,00 | 0,89 |
| Modelo 5 | 0,88 | 0,98 | 0,87 | 0,89 | 0,00 |
El modelo se observó mediante la aplicación RasMol, dando como resultado la proteína presentada en la Figura 5.

Figura 5. Estructura proteica propuesta, modelo 4, para MerB de P. fluorescens. Se observa un monómero, con una estructura estable, que presenta hojas beta, hélices y pequeños giros. Obtenido de RasMol [24]. Al lado derecho se muestra el monómero correspondiente a la estructura usada como molde para el diseño propuesto.
Evaluación y validación de la estructura secundaria para MerB de Pseudomonas fluorescens
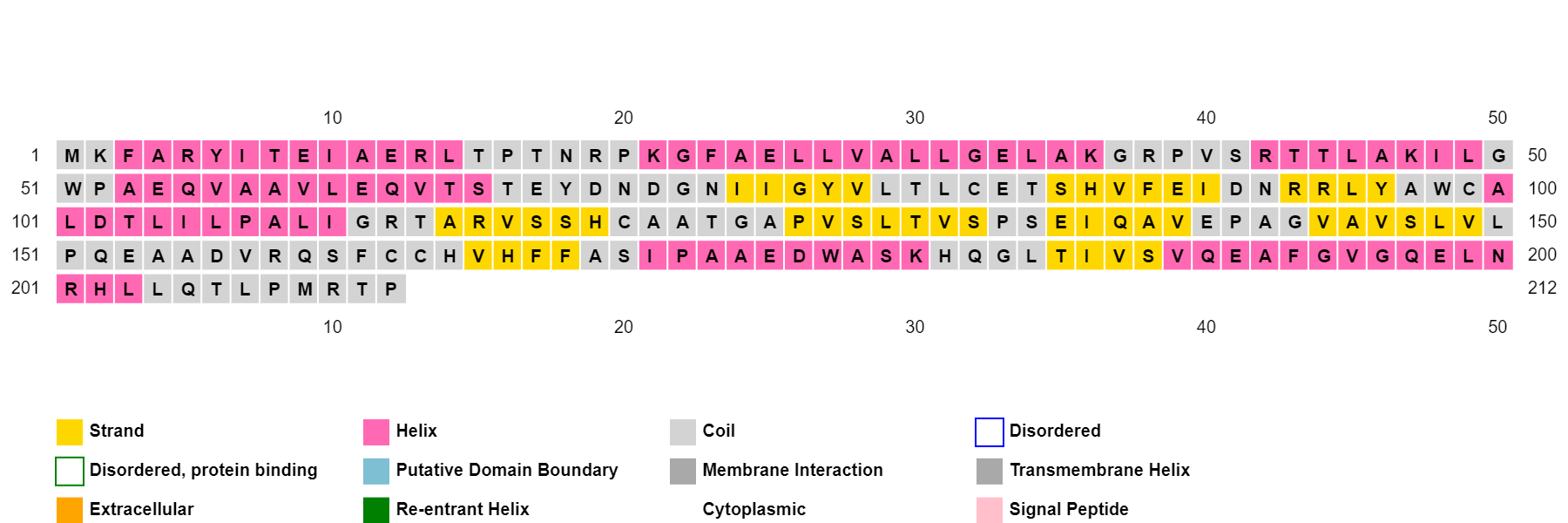
Para el análisis de la estructura secundaria, PSIPRED identifica nueve tramos de aminoácidos con tendencia a formar láminas beta y siete tramos con tendencia a hélices alfa (Figura 6).

Figura 6. Gráfico de secuencia y relación con la estructura secundaria para la enzima de MerB de P. fluorescens, utilizando la aplicación PSIPRED [25]. En amarillo, se identifican los residuos con tendencia a formar hojas beta; en rosa, los residuos con tendencia a formar hélices alfa; y en gris, se muestran los residuos sin tendencia específica a formar una estructura secundaria.
Con la herramienta Biovia se verificó, por medio del gráfico de Ramachandran, la estructura propuesta en la Figura 6, identificando los aminoácidos ubicados en la región de valores permitidos y no permitidos en su estructura secundaria y la distribución de los giros Psi/Phi para el modelo 4, homóloga a P. fluorescens 2 (Figura 7). El diagrama de Ramachandran [33] dado para el modelo 4 de MerB, para P. fluorescens reveló que posee una estructura secundaria con una conformación estable, teniendo el 72,4% de residuos en las regiones más favorables, 21,6% como residuos permitidos, 4,3% en regiones generosamente permitidas y en las zonas no permitidas se encuentra una minoría de residuos del 1,6% diferentes a glicinas, para dar un total de 100% de residuos ubicados en el gráfico.
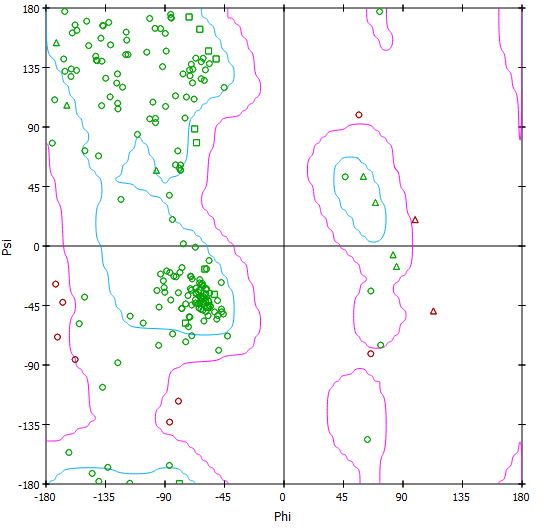
Figura 7. Diagrama de Ramachandran desarrollado por el programa Biovia [26] del modelo 4, en el que se muestra la presencia de cada uno de los aminoácidos de la secuencia en las regiones permitidas y no permitidas para su respectiva estructura secundaria.
Los aminoácidos en el cuadrante superior izquierdo correspondiente a láminas betas paralelas, antiparalelas y giros, logran una mayor cantidad de residuos de hélices alfa con giro hacia la derecha en el cuadrante inferior izquierdo y un porcentaje mínimo de residuos en el cuadrante superior derecho conformados por hélices alfa con giro hacia la izquierda. Además, se identificaron que 185 residuos son diferentes a glicina-prolina, dos son los residuos terminales (exactamente glu y pro), el número de glicinas representado en triángulos es de 11, y el número de prolinas fue es de 11 para un total de 209 aminoácidos.
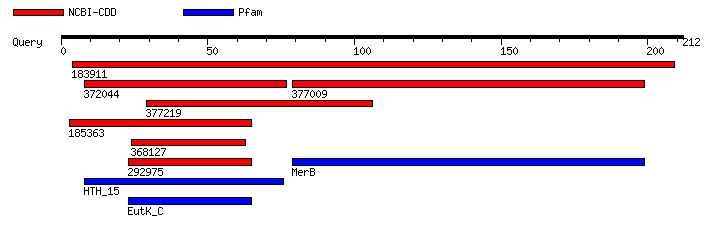
Entre su estructura se encuentran los motivos mostrados en la Figura 8, postulados por NCBI-CDD y Pfam, descritos en las Tablas 4 y 5, con un límite de valor E de 1,0.

Figura 8. Motivos obtenidos a partir de la plataforma MotifSearch [29] mediante las bases de datos NCBI-CDD y Pfam para la secuencia proteica MerB de P. fluorescens.
| Pfam | Posición en la secuencia* | Valor E | Descripción |
| MerB | 79..199 | 2,3e-36 | PF03243, alquilmercurio liasa |
| HTH_15 | 8..76 | 8,4e-23 | PF12324, dominio hélice-vuelta-hélice de la alquilmercurio liasa |
| EutK_C | 23..65 | 0,072 | PF16365, proteína de utilización de etanolamina EutK C-terminal |
(..): representativo de la contracción “al”
Segundo número: aminoácido donde termina su posición en la secuencia proteica de MerB
| NCBI-CDD | Posición en la secuencia* | Valor E | Descripción |
| 183911 | 4..209 | 1e-115 | PRK13239, PRK13239, alquilmercurio liasa MerB |
| 377009 | 79..199 | 1e-38 | pfam03243, MerB, alquilmercurio liasa |
| 372044 | 8..77 | 4e-20 | pfam12324, HTH_15, dominio hélice-vuelta-hélice de la alquilmercurio liasa |
| 377219 | 29..106 | 0,009 | pfam04079, SMC_ScpB, subunidad del complejo de condensación y segregación ScpB |
| 185363 | 3..65 | 0,027 | PRK15466, PRK15466, proteína de microcompartimento de utilización de etanolamina EutK |
| 368127 | 24..63 | 0,28 | pfam04801, Sin_N, región conservada de proteína similar a Sin |
| 292975 | 23..65 | 0,59 | pfam16365, EutK_C, proteína de utilización de etanolamina EutK C-terminal |
(..): representativo de la contracción “al”
Segundo número: aminoácido donde termina su posición en la secuencia proteica de MerB
Entre los motivos encontrados, se escogieron aquellos con un valor E más bajo: el motivo MerB con mayor tamaño en la secuencia con ubicación en el aminoácido 79 al 1.999 con un valor E de 2,3 10-36, seguidamente de HTH_15 con ubicación 8 al 76 y un valor E de 8,4 10-23.
Validación de la estructura tridimensional propuesta para la proteína MerB de Pseudomonas fluorescens
Con el objetivo de validar el modelo 4 postulado como el homólogo de MerB para P. fluorescens, se desarrolló un análisis de calidad general del modelo, utilizando la plataforma Prosa-Web. Como resultado, esta estructura evaluada se calificó con un puntaje Z de −6,5, y se encuentra dentro del rango de puntuaciones típicas de las proteínas nativas de este grupo proteico (Figura 9). Se observa, de igual manera, que la estructura propuesta se encuentra dentro de la sección óptima en la figura del lado izquierdo, verificado por el punto negro. En la figura del lado derecho, se representa la calidad local del modelo por medio de la energía basada en conocimiento; para este análisis se identificó la presencia de picos que corresponden a regiones de paso a través de la membrana de la proteína, lográndose así una estructura totalmente sin partes problemáticas o erróneas en el modelo, evidenciándose en los valores negativos correspondientes.

Figura 9. Datos obtenidos de validación de la estructura tridimensional del modelo 4 mediante la aplicación Prosa-Web [27]: a) Calidad total del modelo a través del puntaje Z (Z-score) y b) Calidad local del modelo por medio de la energía basada en conocimiento.
En seguida, se confirmaron los datos obtenidos de Prosa-Web con la herramienta Verify 3D (Figura 10), mediante un análisis donde el límite máximo esperado es de 100,00%, de los residuos tienen una puntuación 3D/1D promedio > = 0,2. Para el modelo propuesto (modelo 4), se identificó que este presenta el 80% de los aminoácidos que han puntuado con valores > = 0,2 en el perfil 3D/1D, con lo cual se valida que la estructura propuesta, representada en el modelo 4, homóloga para MerB de P. fluorescens, es clasificada de alta calidad y posee coherencia entre la secuencia y la estructura modelada.
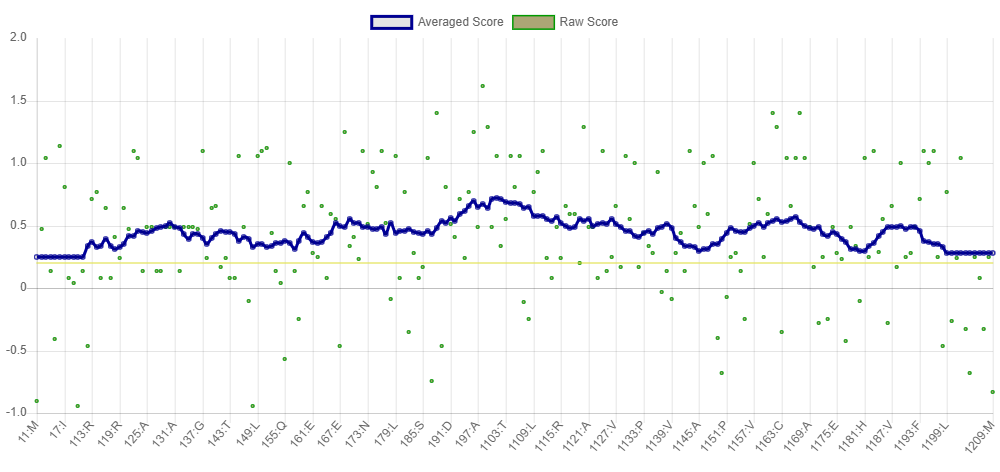
Figura 10. Validación de los datos obtenidos de Prosa-Web mediante la herramienta Verify 3D [34].
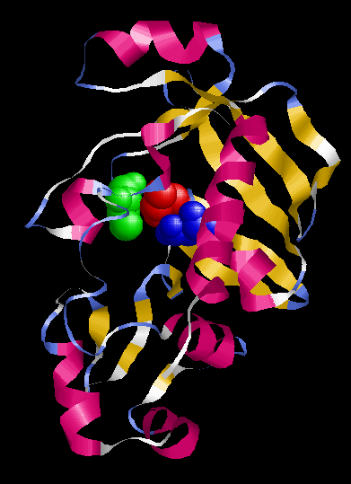
Todos los resultados anteriormente presentados en el documento confirman que el modelo planteado, modelo 4, representa una propuesta bastante sólida de la posible estructura 3D para la proteína MerB de P. fluorescens (Figura 11).

Figura 11. Modelo final propuesto para la proteína MerB de P. fluorescens. En color rosa se muestran las hélices, en amarillo las hojas beta, cintas azules giros y se muestra el sitio activo: en rojo Cys 99, en azul Asp 102 y en verde Cys 162.
El sitio activo se determinó mediante un alineamiento a partir del molde 3FN8 y la secuencia proteica MerB de P. fluorescens por medio de la aplicación del European Bioinformatics Institute [21], T-Coffe [22].
Discusión
En la literatura estudiada, se ha identificado y estructurado la proteína organomercurial liasa (MerB), de la cual no se encuentra una estructura única, puesto que se ha presentado en cada microorganismo con variantes genómicas, encontrándose entre ellos dos arqueas y 11 filos bacterianos [35]. Además, aún no se han reportado estructuras determinadas experimentalmente ni modelamientos de esta proteína, para la bacteria estudiada en el presente artículo, P. fluorescens. Por esta razón, se inició el trabajo desde su secuencia genómica, con el fin de predecir, por medio de la estrategia de modelamiento por homología con técnicas bioinformáticas, un modelo de estructura proteica de esta molécula. Se identificó que la secuencia de MerB de P. fluorescens la constituyen 212aa, de igual manera y con la ayuda del programa Geno3D se estableció que la estructura con mayor similitud a la molécula con la que trabajamos corresponde a la MerB en complejo con mercurio de Escherichia coli (código PDB. 3FN8), ya que fue la que presentó el valor E más bajo (. = 2 × 10-50), por lo que esta fue usada como molde para obtener un modelo presentado.
Mediante el estudio de la estructura proteica de los modelos ya existentes, se identificó la región característica de este tipo de proteínas biorremediadoras de mercurio, definido como el complejo metálico con mercurio. Y es, precisamente, en esta región, su sitio activo, donde surgen las mayores variantes entre las proteínas MerB de cada microorganismo [9]. Sin embargo, es de resaltar que en los estudios de similitud de las proteínas MerB han demostrado ser homólogas entre ellas [14]. En el análisis del alineamiento, todas las secuencias de Pseudomonas sp analizadas para la familia MerB mostraron un 91% de posiciones conservadas.
En nuestro estudio, se identificó que el sitio activo de la familia de las proteínas MerB está conformado, en su mayoría, por un ion mercurio unido a una triada de aminoácidos. En el caso 3FN8 (molde utilizado), posee un ion de mercurio (Hg+2), el cual, por lo regular, forma una triada catalítica con dos residuos de cisteínas (Cys 96 y Cys 159). En su mayoría son conservadas en las secuencias conocidas hasta el momento y que a su vez trabajan con un residuo de ácido aspártico (Asp 99), donde este, por el contrario, es generalmente variante; de esta manera, la indicada en 3FN8 es la variante más presente en las MerB de Pseudomona spp, pues se han encontrado en otras moléculas uniones en este punto con residuos de serina o cisteína, encontrada en pocas variantes [15].
Por otra parte, en estudios anteriores se ha visto que MerB posee un modelo nativo con sitio activo de solo cisteínas, donde evolutivamente se han reportado mutaciones como las ya mencionadas, pero con una actividad catalítica moderadamente más baja; aquella MerB es de solo cisteínas pertenecientes a E. coli, la cual presentó más amplio espectro en compuestos organomercuriales y Hg (II), lo cual no confirmaban que pasaba lo mismo con Pseudomona [36, 37]. Otro reporte que confirma la diferencia de actividad catalítica es la detección de la esencialidad de las cisteínas presentes, siendo la C159 la principal en la catálisis y la C160 con menos efectividad; se recuerda que es en este último aminoácido que P. fluorescens posee un cambio de aminoácido. Por tanto, es en este punto y la razón por la cual es donde se presentan las mutaciones. Sin embargo, la ausencia de cisteína afecta la función catalítica de MerB, puesto que C159/C160 son las más expuestas y flexibles en su estructura, y tienen una función similar al par de cisteínas expuestas en MerA; tener un par de cisteínas exteriores flexibles contribuyen a desplazar los compuestos organomercuriales exógenos, logrando una amplia aceptación del sustrato [14].
En el caso de la molécula modelada, el sitio activo del modelo 4 se identificó mediante un alineamiento por pares a través de la aplicación T-Coffe [22] (Figura 12) entre 3FN8 (molde) y la secuencia de P. fluorescens estudiada.

Figura 12. Alineamiento por pares mediante T-Coffee entre 3FN8 (molde) y la secuencia de P. fluorescens estudiada (con rojo Cys 99, azul Asp 102 y verde Cys 162).
Mediante esta estrategia, se determinó que para el caso de la secuencia a partir de la cual se realizó el modelado, esta presenta exactamente los mismos residuos en las posiciones antes descritas (dos cisteínas y un ácido aspártico). Sin embargo, en el caso del modelo presentado, estas no corresponden exactamente a la ubicación, ya que para su alineamiento fue necesario introducir tres espacios en la secuencia de 3FN8 (molde usado), corriendo exactamente cada aminoácido del sitio activo de P. fluorescenstres aminoácidos a su derecha; por tanto, su sitio activo es Cys 99, Asp 102 y Cys 162.
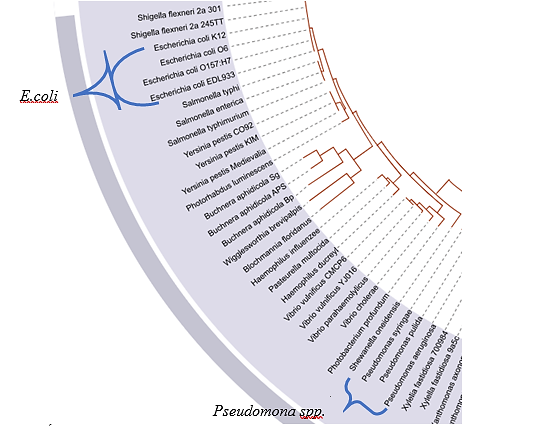
La razón de este cambio de ubicación puede deberse a una razón evolutiva, como se dijo en la presentación de 3FN8; esta proteína fue aislada de E. coli, donde evolutivamente se presenta después de P. fluorescens(Figura 13). Por tanto, se postula que a través del tiempo debido a las mutaciones, E. coli sufrió deleciones en su secuencia, mientras que los genomas de las especies de P. fluorescens aún poseen aquellas regiones como aminoácidos conservados.
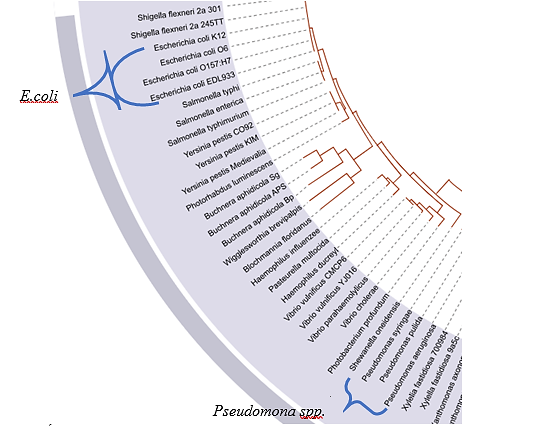
Figura 13. Árbol filogenético de bacterias. Se señala que el género Pseudomona spp apareció evolutivamente primero que E. coli. Tomada y editada de [38].
Además, las mutaciones en este tipo de microorganismos biorremediadores se presentan frecuentemente, demostrándose desde su comienzo, pues los linajes de evolución temprana tienden a presentar solo MerA; pues gracias a las presiones ambientales que ocasionan la evolución, el operón Mer logró reclutar la proteína MerB, la cual hace que los linajes de evolución más recientes presenten ambos precursores. Lo anterior se evidencia desde hace poco tiempo; en la actualidad, hay minoría en homólogos de MerB y pocos estudios en su estructura proteica. Gracias a sus ancestros se logró este cambio, donde MerB tiene su ancestro en un mesófilo y MerA en un termófilo, lo cual hizo que desde ese momento existiera más homólogos mesófilos y no solo termófilos, dando un punto positivo para la biorremediación, logrando que microorganismos como E. coli se encuentre dentro de este tipo de linaje mutando con respecto a P. fluorescens [35].
Pero tal cambio de ubicación no afecta en su mecanismo de acción, donde las cisteínas presentes en el sitio activo poseen la función en la unión del sustrato, la ruptura del enlace carbono-mercurio y la regulación de la liberación de iones de mercurio desde el sitio activo; mientras que su otro aminoácido cumple una función complementaria o si se encuentra una cisteína, esta poseería una función estructural [39], o en este caso si se encuentra el ácido aspártico funciona como un donante de protón [40], ya que después de la escisión química de ligandos nucleofílicos de carbono-mercurio, teniendo alta actividad C-99, es necesario posteriormente la protonólisis del enlace carbono-mercurio, donde se postula que el residuo D-102 desprotonaría a C-99, pues el carboxilato D-102 está más próximo a C-99 (3,6 Å) en forma libre que C-162 (4,3 Å) [36]. Tal mecanismo de acción también se lleva a cabo gracias a la estructura de este mismo sitio activo de MerB, descrito como “sitio activo enterrado”, un proceso llamado “canalización del sustrato”, y es una alternativa a un mecanismo disociativo, ya que en el primer paso donde MerB desprotoniza, se libera el ion mercurio siendo aún más tóxico que el sustrato, el cual es altamente tóxico para las demás células; pero es esta hendidura la que evita tal afectación, pues desplaza al mercurio iónico a MerA para seguir con su reducción y evitar que proceda al citosol [36].
A partir de comparaciones por homología con diferentes secuencias de MerB, se ha establecido, además, que las proteínas MerB no poseen alta especificidad en el reconocimiento del sustrato, puesto que a partir de las secuencias de MerB de bacterias resistentes a organomercuriales aisladas, se evidenció que el ácido aspártico presenta mayor actividad de escindir enlaces C-Hg en comparación con otro residuo como lo es la serina [36]. Según estudios, las proteínas MerB con un residuo de serina están plegadas incorrectamente y no cortan los enlaces carbono-mercurio en las mismas condiciones que MerB [15].
Estructuralmente, el modelo postulado posee una composición estable y se identifica, por medio de la aplicación PSIPRED, una estructura secundaria de nueve tramos de aminoácidos con tendencia a formar láminas beta y siete tramos con tendencia a hélices alfa, unidos a su vez con pequeños giros, los cuales pueden ser confirmados en RasMol (Figura 5); y mediante el gráfico de Ramachandran, se clasifican en una estructura “tradicional”, que favorece su homología hacia las proteínas nativas de su familia y logra una estructura totalmente sin elementos problemáticos o erróneos en el modelo; se evidencia en los puntajes de calidad en ProsaWeb, por lo que se sugiere que el modelo propuesto es coherente entre su secuencia y su modelo tridimensional.
Conclusiones
El modelamiento por homología se reconoce como estrategia útil cuando hay carencia en información experimental reportada en bases de datos y se necesite realizar un ensayo experimental relacionado con la estructura y el funcionamiento de una proteína. Las metodologías existentes para la estructuración de proteínas requieren de alta precisión, ya sea mediante la difracción de rayos X o por resonancia magnética nuclear (RMN), abriendo un campo de investigación en la química computacional. Los resultados del análisis bioinformático de modelamiento por homología indican que la proteína MerB de P. fluorescens es homóloga con la enzima MerB en complejo con mercurio de Escherichia coli 3FN8, el cual es el molde para postular un modelo preciso, el modelo 4. Tales resultados relacionan a la proteína MerB de P. fluorescens como biorremediadora de mercurio, aportando conocimiento en el mecanismo de resistencia a metales más estudiados en la biorremediación. Tales resultados computacionales dan a conocer una plataforma para implementar más estudios y proporcionar referencias sobre el potencial biotecnológico de esta molécula, para el tratamiento de toda fuente contaminada por mercurio a través de sistemas biológicos.
Referencias
[1] I. Bergdahl et al., “Methylmercury and inorganic mercury in serum - Correlation to fish consumption and dental amalgam in a cohort of women born in 1922”, Environmental Research, vol. 77, no. 1, pp. 20-24, 1998, doi: 10.1006/enrs.1997.3820.
[2] G. Genchi, M. Sinicropi, A. Carocci, G. Lauria, and A. Catalano, “Mercury exposure and heart diseases”, International Journal of Environmental Research and Public Health, vol. 14, no. 1. MDPI AG, p. 74, Jan. 12, 2017. doi: 10.3390/ijerph14010074.
[3] G. Compeau and R. Bartha, “Sulfate-reducing bacteria: Principal methylators of mercury in anoxic estuarine sediment”, Applied and Environmental Microbiology, vol. 50, no. 2, pp. 498-502, 1985, doi: 10.1128/aem.50.2.498-502.1985.
[4] B. Debasis and S. Anand, A review on mercury toxicity in foods. 2016.
[5] M. Dua, A. Singh, N. Sethunathan, and A. Johri, “Biotechnology and bioremediation: Successes and limitations”, Applied Microbiology and Biotechnology, vol. 59, no. 2-3. Springer, pp. 143-152, 2002. doi: 10.1007/s00253-002-1024-6.
[6] B. Sharma, A. Dangi, and P. Shukla, “Contemporary enzyme based technologies for bioremediation: A review”, Journal of Environmental Management, vol. 210. Academic Press, pp. 10-22, Mar. 15, 2018. doi: 10.1016/j.jenvman.2017.12.075.
[7] M. Harada et al., “Genomic analysis of Pseudomonas sp. Strain SCT, an iodate-reducing bacterium isolated from marine sediment, reveals a possible use for bioremediation”, G3: Genes, Genomes, Genetics, vol. 9, no. 5, pp. 1321-1329, May 2019, doi: 10.1534/g3.118.200978.
[8] R. Zheng, S. Wu, N. Ma, and C. Sun, “Genetic and physiological adaptations of marine bacterium Pseudomonas stutzeri 273 to mercury stress”, Frontiers in Microbiology, vol. 9, no. APR, Apr. 2018, doi: 10.3389/fmicb.2018.00682.
[9] J. P. Bourdineaud, G. Durn, B. Režun, A. Manceau, and J. Hrenović, “The chemical species of mercury accumulated by Pseudomonas idrijaensis, a bacterium from a rock of the Idrija mercury mine, Slovenia”, Chemosphere, vol. 248, p. 126002, Jun. 2020, doi: 10.1016/j.chemosphere.2020.126002.
[10] A. Nascimento and E. Chartone, “Operon mer: bacterial resistance to mercury and potential for bioremediation of contaminated environments”, Genet. Mol., vol. 2(1), pp. 92-101, 2003.
[11] F. Baldi, M. Gallo, D. Battistel, E. Barbaro, A. Gambaro, and S. Daniele, “A broad mercury resistant strain of Pseudomonas putida secretes pyoverdine under limited iron conditions and high mercury concentrations”, BioMetals, vol. 29, no. 6, pp. 1097-1106, Dec. 2016, doi: 10.1007/s10534-016-9980-y.
[12] T. Begley, A. Walts, and C. Walsh, “Bacterial organomercurial lyase: Overproduction, isolation, and characterization”, Biochemistry, vol. 25, no. 22, pp. 7186-7192, 1986, doi: 10.1021/bi00370a063.
[13] T. Begley, A. Walts, and C. Walsh, “Mechanistic studies of a protonolytic organomercurial cleaving enzyme: Bacterial organomercurial lyase”, Biochemistry, vol. 25, no. 22, pp. 7192-7200, 1986, doi: 10.1021/bi00370a064.
[14] G. Benison et al., “A stable mercury-containing complex of the organomercurial lyase MerB: Catalysis, product release, and direct transfer to MerA”, Biochemistry, vol. 43, no. 26, pp. 8333-8345, Jul. 2004, doi: 10.1021/bi049662h.
[15] R. Karri, R. Das, R. Rai, A. Gopalakrishnan, and G. Roy, “Hg-C bond protonolysis by a functional model of bacterial enzyme organomercurial lyase MerB”, Chemical Communications, vol. 56, no. 65, pp. 9280-9283, Aug. 2020, doi: 10.1039/d0cc02232b.
[16] N. Hamilton and T. Huber, “An introduction to protein contact prediction”, in Methods in Molecular Biology, vol. 453, Humana Press, 2008, pp. 87-104. doi: 10.1007/978-1-60327-429-6_3.
[17] J. Chalarca and D. Gaviria, “Modelamiento in silico por homología de la quinoproteína glucosa deshidrogenasa unida a membrana en Pseudomonas fluorescens implicada en el mecanismo de solubilización de fosfato inorgánico. In silico modeling by homology of membrane-bound quinoprotein g”, Rev. Acad. Colomb. Cienc. Ex. Fis. Nat., vol. 44(173), pp. 1099-1112, 2020, doi: doi.org/10.18257/raccefyn.1154.
[18] A. McPherson, “Protein crystallization”, in Methods in Molecular Biology, vol. 1607, Humana Press Inc., 2017, pp. 17-50. doi: 10.1007/978-1-4939-7000-1_2.
[19] M. Yamamoto, K. Hirata, T. Hikima, Y. Kawano, and G. Ueno, “Protein micro-crystallography with a new micro-beam beamline”, Yakugaku Zasshi, vol. 130, no. 5. Yakugaku Zasshi, pp. 641-648, May 2010. doi: 10.1248/yakushi.130.641.
[20] “UniProt”. https://www.uniprot.org/ (accessed Jan. 02, 2022).
[21] “The European Bioinformatics Institute < EMBL-EBI”. https://www.ebi.ac.uk/ (accessed Jan. 02, 2022).
[22] “T-Coffee Multiple Sequence Alignment Program Using Lalign and Dialign | EBI”. https://www.ebi.ac.uk/Tools/msa/tcoffee/ (accessed Jan. 02, 2022).
[23] I. of B. and P. Chemistry, “Geno3D: Automatic modeling of proteins three-dimensional structure”. https://geno3d-prabi.ibcp.fr/cgi-bin/geno3d_automat.pl?page=/GENO3D/geno3d_home.html (accessed Jan. 02, 2022).
[24] “RasMol and OpenRasMol”. http://www.openrasmol.org/ (accessed Jan. 02, 2022).
[25] “PSIPRED Workbench”. http://bioinf.cs.ucl.ac.uk/psipred/ (accessed Jan. 02, 2022).
[26] “Life Sciences and Material Sciences | BIOVIA - Dassault Systèmes”. https://www.3ds.com/products-services/biovia/ (accessed Jan. 02, 2022).
[27] “ProSA-web - Protein structure analysis”. https://prosa.services.came.sbg.ac.at/prosa.php (accessed Jan. 02, 2022).
[28] D. Eisenberg, R. Lüthy, and J. U. Bowie, “[20] Verify3D: Assessment of protein models with three-dimensional profiles”. Methods in Enzymology, vol. 277, pp. 396-404, Jan. 1997, doi: 10.1016/S0076-6879(97)77022-8.
[29] “MOTIF: Searching protein sequence motifs”. https://www.genome.jp/tools/motif/ (accessed Jan. 02, 2022).
[30] “Phylogeny.fr: Home”. https://www.phylogeny.fr/ (accessed Jan. 02, 2022).
[31] “ExPASy - Compute pI_Mw tool”. https://web.expasy.org/compute_pi/
[32] “RCSB PDB: Homepage”. https://www.rcsb.org/ (accessed Jan. 02, 2022).
[33] G. N. Ramachandran, C. Ramakrishnan, and V. Sasisekharan, “Stereochemistry of polypeptide chain configurations”, Journal of Molecular Biology, vol. 7, no. 1. J Mol Biol, pp. 95-99, 1963. doi: 10.1016/S0022-2836(63)80023-6.
[34] “SAVESv6.0 - Structure validation server”. https://saves.mbi.ucla.edu/ (accessed Jan. 02, 2022).
[35] S. Bhaduri, R. L. Buchanan, and J. G. Phillips, “Expanded response surface model for predicting the effects of temperatures, pH, sodium chloride contents and sodium nitrite concentrations on the growth rate of Yersinia enterocolitica”, Journal of Applied Bacteriology, vol. 79, no. 2, pp. 163-170, Aug. 1995, doi: 10.1111/j.1365-2672.1995.tb00930.x.
[36] J. Lafrance-Vanasse, M. Lefebvre, P. di Lello, J. Sygusch, and J. G. Omichinski, “Crystal structures of the organomercurial lyase MerB in its free and mercury-bound forms Insights into the mechanism of methylmercury degradation”, Journal of Biological Chemistry, vol. 284, no. 2, pp. 938-944, 2008, doi: 10.1074/jbc.M807143200.
[37] Y. Nakahara et al., “Purification and characterization of silkworm hemocytes by flow cytometry”, Dev Comp Immunol, vol. 33, no. 4, pp. 439-448, Apr. 2009, doi: 10.1016/J.DCI.2008.09.005.
[38] I. Letunic and P. Bork, “Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation”, Bioinformatics, vol. 23, no. 1, pp. 127-128, Jan. 2007, doi: 10.1093/bioinformatics/btl529.
[39] J. G. Melnick and G. Parkin, “Cleaving mercury-alkyl bonds: A functional model for mercury detoxification by MerB”, Science (1979), vol. 317, no. 5835, pp. 225-227, Jul. 2007, doi: 10.1126/science.1144314.
[40] H. M. Wahba, M. J. Stevenson, A. Mansour, J. Sygusch, D. E. Wilcox, and J. G. Omichinski, “Structural and biochemical characterization of organotin and organolead compounds binding to the organomercurial lyase MerB provide new insights into its mechanism of carbon-metal bond cleavage”, J Am Chem Soc, vol. 139, no. 2, pp. 910-921, Jan. 2017, doi: 10.1021/Jacs.6b11327.
Notas de autor
kerynd-baronad@unilibre.edu.co
Recibido: 12 de septiembre de 2021; Revisión solicitada: 12 de marzo de 2022; Aceptado: 3 de mayo de 2022
Resumen
El modelamiento in silico ha sido de gran contribución en los procesos proteómicos, desarrollando estructuras de las secuencias proteicas ya existentes, que por motivos de altos costos y las diferentes tecnologías necesarias para el desarrollo de estas metodologías, se encuentran deficientes en el número de modelamientos de proteínas disponibles. Entre aquellas secuencias con carencia de estructura proteica se encuentra la proteína liasa organomercurial (MerB) de Pseudomonas fluorescens, importante en la resistencia al mercurio. En el presente artículo se analizó tanto estructural como funcionalmente la proteína MerB en Pseudomonas fluorescens, utilizando la herramienta de la química estructural “modelamiento por homología” mediante plataformas bioinformáticas, con el fin de obtener un modelo que represente la estructura 3D más precisa y que capturen las mejores variantes estructurales entre todas las posibles conformaciones de las proteínas en la familia. En este trabajo, se desarrolló un método comparativo de la secuencia estudiada con las reportadas en las bases de datos para las proteínas MerB del género Pseudomonas. Se propone un modelo tridimensional para la enzima (MerB) en P. fluorescens, mediante el modelamiento por homología, se muestra la caracterización en la estructura secundaria, terciaria, la caracterización del dominio catalítico y los motivos estructurales presentes.
Palabras clave
bioinformática, modelado por homología, organomercurial liasa, Pseudomonas fluorescens.Abstract
In silico modeling has made a great contribution to proteomic processes, developing structures of the already existing protein sequences, which for reasons of high costs and the different technologies necessary for the development of these methodologies, are deficient in the number of models of available proteins. Among those sequences lacking protein structure is the organomercurial lyase (MerB) protein from Pseudomonas fluorescens, important in mercury resistance. In this article, the MerB protein in Pseudomonas fluorescens was analyzed both structurally and functionally, using the structural chemistry tool “homology modeling” using bioinformatic platforms, in order to obtain a model that represents the most accurate 3D structure and that captures the best structural variants among all the possible conformations of the proteins in the family. In this work, a comparative method of the sequence studied with those reported in the databases for MerB proteins of the genus Pseudomonas was developed. A three-dimensional model for the enzyme (MerB) in P. fluorescens is proposed, through homology modeling, the characterization at the secondary and tertiary structure level, the characterization of the catalytic domain and the structural motifs present is shown.
Keywords
Bioinformatics, homology modelling, organomercurial lyase, Pseudomonas fluorescens .Resumo
A modelagem in silico tem dado um grande contributo para os processos proteómicos, desenvolvendo estruturas de sequências de proteínas já existentes, as quais, pelos elevados custos e pelas diferentes tecnologias necessárias ao desenvolvimento destas metodologias, são deficientes no número de modelos de proteínas disponíveis. Entre as sequências sem estrutura protéica está a proteína organomercurial liase (MerB) de Pseudomonas fluorescens, importante na resistência ao mercúrio. Neste artigo, a proteína MerB em Pseudomonas fluorescens foi analisada estrutural e funcionalmente, usando a ferramenta de química estrutural “modelagem de homologia” usando plataformas de bioinformática, a fim de obter um modelo que represente a estrutura 3D mais precisa e que capture as melhores variantes estruturais. entre todas as conformações possíveis das proteínas da família. Neste trabalho, foi desenvolvido um método comparativo da sequência estudada com aqueles relatados em bancos de dados para proteínas MerB do gênero Pseudomonas. Um modelo tridimensional para a enzima (MerB) em P. fluorescens é proposto, através de modelagem por homologia, a caracterização em nível de estrutura secundária e terciária, a caracterização do domínio catalítico e os motivos estruturais presentes são mostradas.
Palavras-chave
bioinformática, modelagem de homologia, organomercurial Liasa, Pseudomonas fluorescens .Introducción
Desde hace 20.000-30.000 años A. de C., el mercurio ha sido un elemento incluido en la mano de obra del hombre [1], siendo la minería la más reconocida (exactamente en la extracción de oro a base de mercurio) y, a la vez, la actividad que más contaminación provoca por este metal. Por tanto, durante todo este tiempo ha surgido bioacumulación en su mayor reservorio (ríos, lagos y océanos), provocando afectaciones ambientales y en el ser humano [2]. La bioacumulación en los océanos es debida, por lo general, a las bacterias acuáticas anaeróbicas reductoras de sulfato, que transforman el mercurio elemental a metilmercurio por medio de la biometilación de este elemento [3], siendo el estado más contaminante del mercurio; aunque también puede ser transformado por actividades antropogénicas y ser, posteriormente, llevadas a las zonas acuáticas naturales. Afectando de manera directa a organismos acuáticos, y luego a las personas quienes basan su dieta principalmente de productos marinos [4].
En sintonía con la evolución del bienestar ambiental y humano, han surgido diferentes soluciones para este efecto adverso, como lo es la biorremediación, un mecanismo liderado por microorganismos y sus enzimas, para la degradación y transformación de contaminantes a otro estado de oxidación, el cual es menos tóxico para el medioambiente [5]. Entre estos microorganismos se encuentran las bacterias, uno de cuyos géneros es el Pseudomona [6]; las bacterias de este género son organismos Gram negativos de vida libre, con genomas grandes (en promedio 6 Mb), en los cuales tienen la información para adaptarse fácilmente a las condiciones que el ambiente en el que se encuentra le presente. Una de las especies de este género, con una amplia capacidad de adaptación y que ha sido usada comúnmente en procesos de biorremediación, es el Pseudomonas fluorescens. Además, se ha reportado que este microorganismo ha logrado sobrevivir en lugares con alta concentración de mercurio, por lo que la identificación de los elementos genéticos presentes, y que le dan esta característica, se han convertido en un importante tema de estudio [7].
Se ha identificado que los sistemas de resistencia bacteriana a mercurio se agrupan en un operón, conocido como el “operón Mer”, que varía en estructura y consta de genes que codifican proteínas para la regulación (merR, merD), transporte (merT, merP and/or merC, merF) y reducción de mercurio (MerA) [8] y, en algunos casos, un gen adicional, merB, el cual otorga a los microorganismos que lo presentan la capacidad de cortar el enlace carbono-mercurio de organomercuriales como el acetato fenilmercúrico, y generar Hg2+, que luego es desintoxicado como mercurio metálico por la reductasa mercúrica. Se ha identificado la presencia de este gen en la bacteria P. fluorescens, como uno de los elementos importantes de su capacidad de biorremediación de mercurio [9-11]. MerB es la encargada de escindir el enlace carbono-mercurio por medio de una protonólisis produciendo un compuesto alifático o aromático y Hg (II) [12], el cual posteriormente es reducido a Hg (0) por la reductasa específica MerA [13]. MerB muestra una amplia especificidad de sustrato y puede dividir una gran variedad de compuestos organomercuriales que van desde compuestos de alquilo simples como Me-Hg a poliaromáticos y compuestos heterocíclicos como la merbromina [14]. En diversos reportes se evidencia que, para llevar a cabo esta reacción catalítica, los sitios activos de la proteína MerB varían en los diferentes macroorganismos en los que esta ha sido estudiada, como lo es en el dominio bacteria y en las arqueas. Se han identificado mutaciones génicas que en algunos casos producen cambios en la ubicación de algunos aminoácidos; aunque, en general, en la molécula se mantiene una gran similitud. Sin embargo, se ha identificado que estas variaciones ocasionan cambios en la expresión génica y en la estructura proteica [15].
Son estos últimos elementos los que sustentan la importancia de la biología estructural y la química computacional, con el fin de establecer con exactitud la estructura de las proteínas, confiriendo mayor conocimiento de su estructura, funcionalidad, su dinámica, las interacciones con ligandos y otras proteínas [16, 17]. Tal estudio se realiza por medio de herramientas proteómicas, con el propósito de observar las estructuras proteicas como lo es la cristalización con ayuda de la difracción de rayos X o de resonancia magnética, donde la calidad de la imagen estructural final es directamente proporcional a la perfección de las propiedades físicas del espécimen cristalino, que otorga con precisión las coordenadas atómicas, necesitándose cristales de gran tamaño y calidad suficiente para lograr obtener una recopilación de datos precisa. Sin embargo, este tipo de tecnologías son complejas y muy costosas, por lo que para todas las proteínas identificadas no existen datos de estructura generados experimentalmente [18, 19]. Pero más allá de ser un limitante, también impulsó el desarrollo de la química computacional, la cual, hoy en día, se orienta al modelaje y evaluación de proteínas in silico, una variante viable para ahorrar tiempo en el diseño de experimentos con mejores expectativas de éxito y disminuir costos en materiales y reactivos. Entre estas herramientas se encuentra el “modelado por homología”, también conocido como “modelado comparativo” o “modelado basado en plantilla (TBM)”; estas estrategias de modelado de la estructura 3D de una proteína, usan una estructura conocida experimentalmente como molde.
En el presente artículo se plantea estudiar, con el uso de herramientas bioinformáticas, el gen, la secuencia de proteína y dar a conocer una estrategia para modelar y analizar la estructura tridimensional de la enzima MerB en Pseudomonas fluorescens, la cual no ha sido reportada. De igual manera, se plantea llevar a cabo la caracterización de la estructura secundaria, identificación de dominios funcionales, estructurales y la especificación del centro activo de esta importante molécula básica en los procesos de biorremediación de mercurio.
Materiales y métodos
Secuencias de MerB y alineamiento múltiple
Las secuencias de la familia de MerB en varias especies del género Pseudomonas, se obtuvieron de la plataforma de UniProt [20], estas se usaron para la realización de un alineamiento múltiple con la aplicación del European Bioinformatics Institute [21], T-Coffe [22], con el fin de identificar el nivel de diversidad entre ellas y, por tanto, la homología presente.
Modelamiento de la estructura tridimensional de MerB de Pseudomonas fluorescens
Con la secuencia identificada para MerB de Pseudomona fluorescens, se desarrolló un modelamiento por homología con la aplicación Geno3D [23], donde, en primer lugar, se introduce la secuencia y se corre con los valores exactos que propone la aplicación, con el fin de seleccionar el modelo con mejor valor propuesto, donde luego se extrae su respectiva secuencia y para facilitar su visualización se introduce en el software RasMol [24].
Análisis de la estructura secundaria de MerB de Pseudomonas fluorescens
Se predijo la estructura secundaria de MerB mediante la aplicación PSIPRED [25], la cual utiliza un método altamente preciso para la predicción de la estructura secundaria a partir de secuencias de aminoácidos.
Evaluación y validación de la estructura tridimensional de MerB de Pseudomonas fluorescens
Se evaluó la validación de los modelos predichos usando los programas Biovia [26] para verificar la validez de su estructura secundaria; ProSA-Web [27] para su ajuste a estructuras predichas mediante métodos de laboratorio, como la resonancia magnética nuclear o la difracción de rayos X; y Verify 3D [28], con el fin de llevar a cabo la validación de los modelos de los perfiles tridimensionales.
Identificación de dominios proteicos
Los dominios proteicos de la secuencia de MerB en P. fluorescens, se identificaron por medio de la página web de búsqueda MotifSearch [29].
Resultados
Alineamiento por pares y múltiple
A partir de las secuencias de la familia de las proteínas MerB (Tabla 1), se realizó, en la aplicación Phylogeny.fr [30], el alineamiento múltiple y la obtención del respectivo árbol filogenético, identificando en él las secuencias de MerB para P. fluorescens, y se obtuvieron cuatro secuencias en diferentes posiciones de la filogenia, como se muestra en la Figura 1.
Tabla 1.: Código y organismos del género Pseudomona spp aplicados en el estudio filogenético obtenidos de UniProt 20.
Código
Organismo
Q8G9P0
Pseudomonas putida 1 (P_putida1)
A0A1W6QXX0
Pseudomonas putida 2 (P_putida2)
Q9F3W5
Pseudomonas sp (P_sp1)
O07303
Pseudomonas sp (P_sp2)
Q9Z3Y8
Pseudomonas sp (P_sp3)
Q9F3W6
Pseudomonas sp (P_sp4)
U2A0X4
Pseudomonas sp (P_sp5)
A0A2N7XWW6
Pseudomonas sp (P_sp6)
Q0E6C2|
Pseudomonas aeruginosa (P_aeruginosa1)
A0A2N7XXX7|
Pseudomonas sp (P_sp7)
A0A212BER9|
Pseudomonas sp (P_sp8)
A0A3M5Z239|
Pseudomonas coronafaciens (P_coronafaciens1)
A0A1G8QSD9|
Pseudomonas abietaniphila (P_abietaniphila1)
A0A5C5R0G9|
Pseudomonas mandelii (P_mandelii1)
A0A5P9WB02|
Pseudomonas aeruginosa (P_aeruginosa2)
A0A1G5PHN0|
Pseudomonas psychrotolerans (P_psychrotolerans1)
A0A2S7FMI2|
Pseudomonas oleovorans (P_oleovorans1)
A0A2N7XWV5
Pseudomonas sp P_sp9
A0A1V0M5B3|
Pseudomonas aeruginosa (P_aeruginosa3)
A0A5B0BVV8|
Pseudomonas sp (P_sp10)
A0A2N7XX29|
Pseudomonas sp (P_sp11)
C0KJV0|
Pseudomonas aeruginosa (P_aeruginosa4)
E1V302|
Pseudomonas plecoglossicida (P_plecoglossicida1)
A4V7U8|
Pseudomonas fluorescens (P_fluorescens1)
S2FX69|
Pseudomonas sp (P_sp12)
A0A5C5N7Q8|
Pseudomonas rhodesiae (P_rhodesiae1)
A0A5E7MFD1|
Pseudomonas fluorescens (P_fluorescens2)
A0A5M8ECB2|
Pseudomonas veronii (P_veronii1)
A0A0R3BTV1|
Pseudomonas lactis (P_lactis1)
A0A419UJ99|
Pseudomonas synxantha (P_synxantha1)
A0A2G0W2S4|
Pseudomonas sp (P_sp13)
A0A127HQV2|
Pseudomonas azotoformans (P_azotoformans)
A0A0R3B0W4|
Pseudomonas lactis (P_lactis2)
A0A3M5ZUV3|
Pseudomonas coronafaciens (P_coronafaciens2)
A0A5E7A816|
Pseudomonas fluorescens (P_fluorescens3)
Q70MS1|
Pseudomonas fluorescens (P_fluorescens4)
E1V304|
Pseudomonas sp (P_sp14)
A0A1E4X3J7|
Pseudomonas sp (P_sp15)
A0A1T1HZC6|
Pseudomonas sp (P_sp16)
E1V303|
Pseudomonas putida (P_putida3)
A0A6B7Q571|
Pseudomonas putida (P_putida4)
A0A1H0ISK0
Pseudomonas poael (P_poae1)
A0A0P9GCM1
Pseudomonas syringae (P_syringae1)
A0A3S4N2X6|
Pseudomonas putrefaciens (S_putrefaciens2)
Figura 1. Árbol filogenético producto del alineamiento en Phylogeny.fr [30] de las secuencias reportadas de Pseudomonas para MerB (se resaltan en color azul los aislados de P. fluorescens, en rojo se observan los valores de Bootstrap, con 500 repeticiones, verificando la estabilidad de cada agrupamiento).
A partir del árbol filogenético se observan las diferentes secuencias de MerB de Pseudomona fluorescens, y se identifica la diversidad entre ellas, especialmente la secuencia “P. fluorescens 2”, que tiene mayor similitud con P. syringae, y es diferente a los demás representantes de su especie en, al menos, el 5%. Teniendo en cuenta este nivel de diferencia con las otras secuencias correspondientes en la especie, se escogió la secuencia representativa para MerB de P. fluorescens para el análisis del modelamiento por homología.
Caracterización fisicoquímica de la enzima MerB de P. fluorescens
Partiendo de la secuencia proteica seleccionada de A0A5E7MFD1 para MerB identificada en P. fluorescens(212aa), se verificó por medio de la plataforma Expasy [31] que su peso molecular es de 23139,4 Da y su punto isoeléctrico es de 5,86. Mediante la plataforma Uniprot [20] se estableció su código EC 4.99.1.2, correspondiente a una liasa de alquilmercurio, para la cual su reacción enzimática se presenta en la Figura 2.
Figura 2. Reacción enzimática de MerB identificada para P. fluorescens,dando lugar a una reacción catabólica teniendo como sustratos un alquilmercurio y un protón de hidrógeno, que produce un alcano y un ion mercurio. Tomada de Uniprot [20].
Modelamiento de la estructura tridimensional de la proteína MerB de P. fluorescens
Con los resultados suministrados por Geno3D, se estableció que la mejor secuencia para el modelamiento por homología es la proteína MerB en complejo con mercurio de Escherichia coli 3FN8 (Figura 3). Esta secuencia presentó mayor similitud a la secuencia MerB de P. fluorescens 2 con un valor E de 2 × 10-50 y una desviación media de 0,48 Å.
Figura 3. Modelo 3FN8 computacional generado a partir del estudio por homología para MerB en P. fluorescens a través de RasMol [24], observándose una proteína dimérica, cada cadena con un centro activo de mercurio (esferas rosa) y glicerol (estructura atómica), unidos covalentemente a residuos de aminoácidos.
Cada centro activo se logra (observar en la Figura 4) formado por un complejo de mercurio, el cual es clasificado como un complejo metálico, donde se sitúa un ion de mercurio (Hg+2) unido covalentemente mediante enlace metálico a dos cisteínas (Cys 96 y Cys 159) y a un ácido aspártico (Asp 99) y por medio de un puente de hidrógeno a un glicerol (Gol 222).
Figura 4. Ilustración computacional del complejo metálico de mercurio para MerB de E. coli 3FN8. Tomada de RCSB PDB Protein Data Bank [32].
Tomando a 3FN8 como molde para el diseño de MerB de P. fluorescens, se generaron cinco modelos (Tabla 2), donde los valores de correlación entre los cinco modelos evaluados se determinaron mediante desviación de la raíz cuadrada media (DRCM) con valores en ángstroms (Tabla 3).
Tabla 2.: Comparación de la calidad de los modelos producidos por el servidor Geno3D. Todos los modelos utilizaron el mismo molde 3FN8. Cada columna representa: los valores de energía en Kcal/mol, el porcentaje de residuos que se encuentran en la región nuclear y el porcentaje de residuos no permitidos en el diagrama de Ramachandran. Los valores de energía indican la fiabilidad y la precisión de la estructura.
Nombre del modelo
Molde
Energía del modelo Kcal/mol
Porcentaje de residuos en la región núcleo del diagrama de Ramachandran
Porcentaje de residuos no permitidos en el diagrama de Ramachandran
Modelo 1
3FN8
-7.860,73
70,3
3,8
Modelo 2
3FN8
4.400,65
76,8
2,2
Modelo 3
3FN8
3.824,18
73,0
1,1
Modelo 4
3FN8
-7.859,05
72,4
1,6
Modelo 5
3FN8
-7.970,61
73,5
1,1
A partir de los valores obtenidos, se escogió el mejor modelo, el cual corresponde al valor negativo más bajo: el modelo 4; además, tiene porcentajes permitidos en el gráfico de Ramachandran.
Tabla 3.: Ajuste estructural entre los modelos DRCM en Å Desviación media 0882754 Å.
Modelo 1
Modelo 2
Modelo 3
Modelo 4
Modelo 5
Modelo 1
0,00
0,77
0,88
0,81
0,88
Modelo 2
0,77
0,00
0,95
0,92
0,98
Modelo 3
0,88
0,95
0,00
0,88
0,87
Modelo 4
0,81
0,92
0,88
0,00
0,89
Modelo 5
0,88
0,98
0,87
0,89
0,00
El modelo se observó mediante la aplicación RasMol, dando como resultado la proteína presentada en la Figura 5.
Figura 5. Estructura proteica propuesta, modelo 4, para MerB de P. fluorescens. Se observa un monómero, con una estructura estable, que presenta hojas beta, hélices y pequeños giros. Obtenido de RasMol [24]. Al lado derecho se muestra el monómero correspondiente a la estructura usada como molde para el diseño propuesto.
Evaluación y validación de la estructura secundaria para MerB de Pseudomonas fluorescens
Para el análisis de la estructura secundaria, PSIPRED identifica nueve tramos de aminoácidos con tendencia a formar láminas beta y siete tramos con tendencia a hélices alfa (Figura 6).
Figura 6. Gráfico de secuencia y relación con la estructura secundaria para la enzima de MerB de P. fluorescens, utilizando la aplicación PSIPRED [25]. En amarillo, se identifican los residuos con tendencia a formar hojas beta; en rosa, los residuos con tendencia a formar hélices alfa; y en gris, se muestran los residuos sin tendencia específica a formar una estructura secundaria.
Con la herramienta Biovia se verificó, por medio del gráfico de Ramachandran, la estructura propuesta en la Figura 6, identificando los aminoácidos ubicados en la región de valores permitidos y no permitidos en su estructura secundaria y la distribución de los giros Psi/Phi para el modelo 4, homóloga a P. fluorescens 2 (Figura 7). El diagrama de Ramachandran [33] dado para el modelo 4 de MerB, para P. fluorescens reveló que posee una estructura secundaria con una conformación estable, teniendo el 72,4% de residuos en las regiones más favorables, 21,6% como residuos permitidos, 4,3% en regiones generosamente permitidas y en las zonas no permitidas se encuentra una minoría de residuos del 1,6% diferentes a glicinas, para dar un total de 100% de residuos ubicados en el gráfico.
Figura 7. Diagrama de Ramachandran desarrollado por el programa Biovia [26] del modelo 4, en el que se muestra la presencia de cada uno de los aminoácidos de la secuencia en las regiones permitidas y no permitidas para su respectiva estructura secundaria.
Los aminoácidos en el cuadrante superior izquierdo correspondiente a láminas betas paralelas, antiparalelas y giros, logran una mayor cantidad de residuos de hélices alfa con giro hacia la derecha en el cuadrante inferior izquierdo y un porcentaje mínimo de residuos en el cuadrante superior derecho conformados por hélices alfa con giro hacia la izquierda. Además, se identificaron que 185 residuos son diferentes a glicina-prolina, dos son los residuos terminales (exactamente glu y pro), el número de glicinas representado en triángulos es de 11, y el número de prolinas fue es de 11 para un total de 209 aminoácidos.
Entre su estructura se encuentran los motivos mostrados en la Figura 8, postulados por NCBI-CDD y Pfam, descritos en las Tablas 4 y 5, con un límite de valor E de 1,0.
Figura 8. Motivos obtenidos a partir de la plataforma MotifSearch [29] mediante las bases de datos NCBI-CDD y Pfam para la secuencia proteica MerB de P. fluorescens.
*Primer número: aminoácido donde inicia su posición en la secuencia proteica de MerB (..): representativo de la contracción “al” Segundo número: aminoácido donde termina su posición en la secuencia proteica de MerBTabla 4.: Descripción de motivos resultantes del análisis en MotifSearch [29] mediante la base de datos Pfam para la secuencia proteica MerB de P fluorescens.
Pfam
Posición en la secuencia*
Valor E
Descripción
MerB
79..199
2,3e-36
PF03243, alquilmercurio liasa
HTH_15
8..76
8,4e-23
PF12324, dominio hélice-vuelta-hélice de la alquilmercurio liasa
EutK_C
23..65
0,072
PF16365, proteína de utilización de etanolamina EutK C-terminal
*Primer número: aminoácido donde inicia su posición en la secuencia proteica de MerB (..): representativo de la contracción “al” Segundo número: aminoácido donde termina su posición en la secuencia proteica de MerBTabla 5.: Descripción de motivos resultantes del análisis en MotifSearch 29 mediante la base de datos NCBICDD para la secuencia proteica MerB de P fluorescens.
NCBI-CDD
Posición en la secuencia*
Valor E
Descripción
183911
4..209
1e-115
PRK13239, PRK13239, alquilmercurio liasa MerB
377009
79..199
1e-38
pfam03243, MerB, alquilmercurio liasa
372044
8..77
4e-20
pfam12324, HTH_15, dominio hélice-vuelta-hélice de la alquilmercurio liasa
377219
29..106
0,009
pfam04079, SMC_ScpB, subunidad del complejo de condensación y segregación ScpB
185363
3..65
0,027
PRK15466, PRK15466, proteína de microcompartimento de utilización de etanolamina EutK
368127
24..63
0,28
pfam04801, Sin_N, región conservada de proteína similar a Sin
292975
23..65
0,59
pfam16365, EutK_C, proteína de utilización de etanolamina EutK C-terminal
Entre los motivos encontrados, se escogieron aquellos con un valor E más bajo: el motivo MerB con mayor tamaño en la secuencia con ubicación en el aminoácido 79 al 1.999 con un valor E de 2,3 10-36, seguidamente de HTH_15 con ubicación 8 al 76 y un valor E de 8,4 10-23.
Validación de la estructura tridimensional propuesta para la proteína MerB de Pseudomonas fluorescens
Con el objetivo de validar el modelo 4 postulado como el homólogo de MerB para P. fluorescens, se desarrolló un análisis de calidad general del modelo, utilizando la plataforma Prosa-Web. Como resultado, esta estructura evaluada se calificó con un puntaje Z de −6,5, y se encuentra dentro del rango de puntuaciones típicas de las proteínas nativas de este grupo proteico (Figura 9). Se observa, de igual manera, que la estructura propuesta se encuentra dentro de la sección óptima en la figura del lado izquierdo, verificado por el punto negro. En la figura del lado derecho, se representa la calidad local del modelo por medio de la energía basada en conocimiento; para este análisis se identificó la presencia de picos que corresponden a regiones de paso a través de la membrana de la proteína, lográndose así una estructura totalmente sin partes problemáticas o erróneas en el modelo, evidenciándose en los valores negativos correspondientes.
Figura 9. Datos obtenidos de validación de la estructura tridimensional del modelo 4 mediante la aplicación Prosa-Web [27]: a) Calidad total del modelo a través del puntaje Z (Z-score) y b) Calidad local del modelo por medio de la energía basada en conocimiento.
En seguida, se confirmaron los datos obtenidos de Prosa-Web con la herramienta Verify 3D (Figura 10), mediante un análisis donde el límite máximo esperado es de 100,00%, de los residuos tienen una puntuación 3D/1D promedio > = 0,2. Para el modelo propuesto (modelo 4), se identificó que este presenta el 80% de los aminoácidos que han puntuado con valores > = 0,2 en el perfil 3D/1D, con lo cual se valida que la estructura propuesta, representada en el modelo 4, homóloga para MerB de P. fluorescens, es clasificada de alta calidad y posee coherencia entre la secuencia y la estructura modelada.
Figura 10. Validación de los datos obtenidos de Prosa-Web mediante la herramienta Verify 3D [34].
Todos los resultados anteriormente presentados en el documento confirman que el modelo planteado, modelo 4, representa una propuesta bastante sólida de la posible estructura 3D para la proteína MerB de P. fluorescens (Figura 11).
Figura 11. Modelo final propuesto para la proteína MerB de P. fluorescens. En color rosa se muestran las hélices, en amarillo las hojas beta, cintas azules giros y se muestra el sitio activo: en rojo Cys 99, en azul Asp 102 y en verde Cys 162.
El sitio activo se determinó mediante un alineamiento a partir del molde 3FN8 y la secuencia proteica MerB de P. fluorescens por medio de la aplicación del European Bioinformatics Institute [21], T-Coffe [22].
Discusión
En la literatura estudiada, se ha identificado y estructurado la proteína organomercurial liasa (MerB), de la cual no se encuentra una estructura única, puesto que se ha presentado en cada microorganismo con variantes genómicas, encontrándose entre ellos dos arqueas y 11 filos bacterianos [35]. Además, aún no se han reportado estructuras determinadas experimentalmente ni modelamientos de esta proteína, para la bacteria estudiada en el presente artículo, P. fluorescens. Por esta razón, se inició el trabajo desde su secuencia genómica, con el fin de predecir, por medio de la estrategia de modelamiento por homología con técnicas bioinformáticas, un modelo de estructura proteica de esta molécula. Se identificó que la secuencia de MerB de P. fluorescens la constituyen 212aa, de igual manera y con la ayuda del programa Geno3D se estableció que la estructura con mayor similitud a la molécula con la que trabajamos corresponde a la MerB en complejo con mercurio de Escherichia coli (código PDB. 3FN8), ya que fue la que presentó el valor E más bajo (. = 2 × 10-50), por lo que esta fue usada como molde para obtener un modelo presentado.
Mediante el estudio de la estructura proteica de los modelos ya existentes, se identificó la región característica de este tipo de proteínas biorremediadoras de mercurio, definido como el complejo metálico con mercurio. Y es, precisamente, en esta región, su sitio activo, donde surgen las mayores variantes entre las proteínas MerB de cada microorganismo [9]. Sin embargo, es de resaltar que en los estudios de similitud de las proteínas MerB han demostrado ser homólogas entre ellas [14]. En el análisis del alineamiento, todas las secuencias de Pseudomonas sp analizadas para la familia MerB mostraron un 91% de posiciones conservadas.
En nuestro estudio, se identificó que el sitio activo de la familia de las proteínas MerB está conformado, en su mayoría, por un ion mercurio unido a una triada de aminoácidos. En el caso 3FN8 (molde utilizado), posee un ion de mercurio (Hg+2), el cual, por lo regular, forma una triada catalítica con dos residuos de cisteínas (Cys 96 y Cys 159). En su mayoría son conservadas en las secuencias conocidas hasta el momento y que a su vez trabajan con un residuo de ácido aspártico (Asp 99), donde este, por el contrario, es generalmente variante; de esta manera, la indicada en 3FN8 es la variante más presente en las MerB de Pseudomona spp, pues se han encontrado en otras moléculas uniones en este punto con residuos de serina o cisteína, encontrada en pocas variantes [15].
Por otra parte, en estudios anteriores se ha visto que MerB posee un modelo nativo con sitio activo de solo cisteínas, donde evolutivamente se han reportado mutaciones como las ya mencionadas, pero con una actividad catalítica moderadamente más baja; aquella MerB es de solo cisteínas pertenecientes a E. coli, la cual presentó más amplio espectro en compuestos organomercuriales y Hg (II), lo cual no confirmaban que pasaba lo mismo con Pseudomona [36, 37]. Otro reporte que confirma la diferencia de actividad catalítica es la detección de la esencialidad de las cisteínas presentes, siendo la C159 la principal en la catálisis y la C160 con menos efectividad; se recuerda que es en este último aminoácido que P. fluorescens posee un cambio de aminoácido. Por tanto, es en este punto y la razón por la cual es donde se presentan las mutaciones. Sin embargo, la ausencia de cisteína afecta la función catalítica de MerB, puesto que C159/C160 son las más expuestas y flexibles en su estructura, y tienen una función similar al par de cisteínas expuestas en MerA; tener un par de cisteínas exteriores flexibles contribuyen a desplazar los compuestos organomercuriales exógenos, logrando una amplia aceptación del sustrato [14].
En el caso de la molécula modelada, el sitio activo del modelo 4 se identificó mediante un alineamiento por pares a través de la aplicación T-Coffe [22] (Figura 12) entre 3FN8 (molde) y la secuencia de P. fluorescens estudiada.
Figura 12. Alineamiento por pares mediante T-Coffee entre 3FN8 (molde) y la secuencia de P. fluorescens estudiada (con rojo Cys 99, azul Asp 102 y verde Cys 162).
Mediante esta estrategia, se determinó que para el caso de la secuencia a partir de la cual se realizó el modelado, esta presenta exactamente los mismos residuos en las posiciones antes descritas (dos cisteínas y un ácido aspártico). Sin embargo, en el caso del modelo presentado, estas no corresponden exactamente a la ubicación, ya que para su alineamiento fue necesario introducir tres espacios en la secuencia de 3FN8 (molde usado), corriendo exactamente cada aminoácido del sitio activo de P. fluorescenstres aminoácidos a su derecha; por tanto, su sitio activo es Cys 99, Asp 102 y Cys 162.
La razón de este cambio de ubicación puede deberse a una razón evolutiva, como se dijo en la presentación de 3FN8; esta proteína fue aislada de E. coli, donde evolutivamente se presenta después de P. fluorescens(Figura 13). Por tanto, se postula que a través del tiempo debido a las mutaciones, E. coli sufrió deleciones en su secuencia, mientras que los genomas de las especies de P. fluorescens aún poseen aquellas regiones como aminoácidos conservados.
Figura 13. Árbol filogenético de bacterias. Se señala que el género Pseudomona spp apareció evolutivamente primero que E. coli. Tomada y editada de [38].
Además, las mutaciones en este tipo de microorganismos biorremediadores se presentan frecuentemente, demostrándose desde su comienzo, pues los linajes de evolución temprana tienden a presentar solo MerA; pues gracias a las presiones ambientales que ocasionan la evolución, el operón Mer logró reclutar la proteína MerB, la cual hace que los linajes de evolución más recientes presenten ambos precursores. Lo anterior se evidencia desde hace poco tiempo; en la actualidad, hay minoría en homólogos de MerB y pocos estudios en su estructura proteica. Gracias a sus ancestros se logró este cambio, donde MerB tiene su ancestro en un mesófilo y MerA en un termófilo, lo cual hizo que desde ese momento existiera más homólogos mesófilos y no solo termófilos, dando un punto positivo para la biorremediación, logrando que microorganismos como E. coli se encuentre dentro de este tipo de linaje mutando con respecto a P. fluorescens [35].
Pero tal cambio de ubicación no afecta en su mecanismo de acción, donde las cisteínas presentes en el sitio activo poseen la función en la unión del sustrato, la ruptura del enlace carbono-mercurio y la regulación de la liberación de iones de mercurio desde el sitio activo; mientras que su otro aminoácido cumple una función complementaria o si se encuentra una cisteína, esta poseería una función estructural [39], o en este caso si se encuentra el ácido aspártico funciona como un donante de protón [40], ya que después de la escisión química de ligandos nucleofílicos de carbono-mercurio, teniendo alta actividad C-99, es necesario posteriormente la protonólisis del enlace carbono-mercurio, donde se postula que el residuo D-102 desprotonaría a C-99, pues el carboxilato D-102 está más próximo a C-99 (3,6 Å) en forma libre que C-162 (4,3 Å) [36]. Tal mecanismo de acción también se lleva a cabo gracias a la estructura de este mismo sitio activo de MerB, descrito como “sitio activo enterrado”, un proceso llamado “canalización del sustrato”, y es una alternativa a un mecanismo disociativo, ya que en el primer paso donde MerB desprotoniza, se libera el ion mercurio siendo aún más tóxico que el sustrato, el cual es altamente tóxico para las demás células; pero es esta hendidura la que evita tal afectación, pues desplaza al mercurio iónico a MerA para seguir con su reducción y evitar que proceda al citosol [36].
A partir de comparaciones por homología con diferentes secuencias de MerB, se ha establecido, además, que las proteínas MerB no poseen alta especificidad en el reconocimiento del sustrato, puesto que a partir de las secuencias de MerB de bacterias resistentes a organomercuriales aisladas, se evidenció que el ácido aspártico presenta mayor actividad de escindir enlaces C-Hg en comparación con otro residuo como lo es la serina [36]. Según estudios, las proteínas MerB con un residuo de serina están plegadas incorrectamente y no cortan los enlaces carbono-mercurio en las mismas condiciones que MerB [15].
Estructuralmente, el modelo postulado posee una composición estable y se identifica, por medio de la aplicación PSIPRED, una estructura secundaria de nueve tramos de aminoácidos con tendencia a formar láminas beta y siete tramos con tendencia a hélices alfa, unidos a su vez con pequeños giros, los cuales pueden ser confirmados en RasMol (Figura 5); y mediante el gráfico de Ramachandran, se clasifican en una estructura “tradicional”, que favorece su homología hacia las proteínas nativas de su familia y logra una estructura totalmente sin elementos problemáticos o erróneos en el modelo; se evidencia en los puntajes de calidad en ProsaWeb, por lo que se sugiere que el modelo propuesto es coherente entre su secuencia y su modelo tridimensional.
Conclusiones
El modelamiento por homología se reconoce como estrategia útil cuando hay carencia en información experimental reportada en bases de datos y se necesite realizar un ensayo experimental relacionado con la estructura y el funcionamiento de una proteína. Las metodologías existentes para la estructuración de proteínas requieren de alta precisión, ya sea mediante la difracción de rayos X o por resonancia magnética nuclear (RMN), abriendo un campo de investigación en la química computacional. Los resultados del análisis bioinformático de modelamiento por homología indican que la proteína MerB de P. fluorescens es homóloga con la enzima MerB en complejo con mercurio de Escherichia coli 3FN8, el cual es el molde para postular un modelo preciso, el modelo 4. Tales resultados relacionan a la proteína MerB de P. fluorescens como biorremediadora de mercurio, aportando conocimiento en el mecanismo de resistencia a metales más estudiados en la biorremediación. Tales resultados computacionales dan a conocer una plataforma para implementar más estudios y proporcionar referencias sobre el potencial biotecnológico de esta molécula, para el tratamiento de toda fuente contaminada por mercurio a través de sistemas biológicos.
Referencias
Referencias
I. Bergdahl et al., “Methylmercury and inorganic mercury in serum - Correlation to fish consumption and dental amalgam in a cohort of women born in 1922”, Environmental Research, vol. 77, no. 1, pp. 20-24, 1998, doi: 10.1006/enrs.1997.3820.
G. Genchi, M. Sinicropi, A. Carocci, G. Lauria, and A. Catalano, “Mercury exposure and heart diseases”, International Journal of Environmental Research and Public Health, vol. 14, no. 1. MDPI AG, p. 74, Jan. 12, 2017. doi: 10.3390/ijerph14010074.
G. Compeau and R. Bartha, “Sulfate-reducing bacteria: Principal methylators of mercury in anoxic estuarine sediment”, Applied and Environmental Microbiology, vol. 50, no. 2, pp. 498-502, 1985, doi: 10.1128/aem.50.2.498-502.1985.
B. Debasis and S. Anand, A review on mercury toxicity in foods. 2016.
M. Dua, A. Singh, N. Sethunathan, and A. Johri, “Biotechnology and bioremediation: Successes and limitations”, Applied Microbiology and Biotechnology, vol. 59, no. 2-3. Springer, pp. 143-152, 2002. doi: 10.1007/s00253-002-1024-6.
B. Sharma, A. Dangi, and P. Shukla, “Contemporary enzyme based technologies for bioremediation: A review”, Journal of Environmental Management, vol. 210. Academic Press, pp. 10-22, Mar. 15, 2018. doi: 10.1016/j.jenvman.2017.12.075.
M. Harada et al., “Genomic analysis of Pseudomonas sp. Strain SCT, an iodate-reducing bacterium isolated from marine sediment, reveals a possible use for bioremediation”, G3: Genes, Genomes, Genetics, vol. 9, no. 5, pp. 1321-1329, May 2019, doi: 10.1534/g3.118.200978.
R. Zheng, S. Wu, N. Ma, and C. Sun, “Genetic and physiological adaptations of marine bacterium Pseudomonas stutzeri 273 to mercury stress”, Frontiers in Microbiology, vol. 9, no. APR, Apr. 2018, doi: 10.3389/fmicb.2018.00682.
J. P. Bourdineaud, G. Durn, B. Režun, A. Manceau, and J. Hrenović, “The chemical species of mercury accumulated by Pseudomonas idrijaensis, a bacterium from a rock of the Idrija mercury mine, Slovenia”, Chemosphere, vol. 248, p. 126002, Jun. 2020, doi: 10.1016/j.chemosphere.2020.126002.
A. Nascimento and E. Chartone, “Operon mer: bacterial resistance to mercury and potential for bioremediation of contaminated environments”, Genet. Mol., vol. 2(1), pp. 92-101, 2003.
F. Baldi, M. Gallo, D. Battistel, E. Barbaro, A. Gambaro, and S. Daniele, “A broad mercury resistant strain of Pseudomonas putida secretes pyoverdine under limited iron conditions and high mercury concentrations”, BioMetals, vol. 29, no. 6, pp. 1097-1106, Dec. 2016, doi: 10.1007/s10534-016-9980-y.
T. Begley, A. Walts, and C. Walsh, “Bacterial organomercurial lyase: Overproduction, isolation, and characterization”, Biochemistry, vol. 25, no. 22, pp. 7186-7192, 1986, doi: 10.1021/bi00370a063.
T. Begley, A. Walts, and C. Walsh, “Mechanistic studies of a protonolytic organomercurial cleaving enzyme: Bacterial organomercurial lyase”, Biochemistry, vol. 25, no. 22, pp. 7192-7200, 1986, doi: 10.1021/bi00370a064.
G. Benison et al., “A stable mercury-containing complex of the organomercurial lyase MerB: Catalysis, product release, and direct transfer to MerA”, Biochemistry, vol. 43, no. 26, pp. 8333-8345, Jul. 2004, doi: 10.1021/bi049662h.
R. Karri, R. Das, R. Rai, A. Gopalakrishnan, and G. Roy, “Hg-C bond protonolysis by a functional model of bacterial enzyme organomercurial lyase MerB”, Chemical Communications, vol. 56, no. 65, pp. 9280-9283, Aug. 2020, doi: 10.1039/d0cc02232b.
N. Hamilton and T. Huber, “An introduction to protein contact prediction”, in Methods in Molecular Biology, vol. 453, Humana Press, 2008, pp. 87-104. doi: 10.1007/978-1-60327-429-6_3.
J. Chalarca and D. Gaviria, “Modelamiento in silico por homología de la quinoproteína glucosa deshidrogenasa unida a membrana en Pseudomonas fluorescens implicada en el mecanismo de solubilización de fosfato inorgánico. In silico modeling by homology of membrane-bound quinoprotein g”, Rev. Acad. Colomb. Cienc. Ex. Fis. Nat., vol. 44(173), pp. 1099-1112, 2020, doi: doi.org/10.18257/raccefyn.1154.
A. McPherson, “Protein crystallization”, in Methods in Molecular Biology, vol. 1607, Humana Press Inc., 2017, pp. 17-50. doi: 10.1007/978-1-4939-7000-1_2.
M. Yamamoto, K. Hirata, T. Hikima, Y. Kawano, and G. Ueno, “Protein micro-crystallography with a new micro-beam beamline”, Yakugaku Zasshi, vol. 130, no. 5. Yakugaku Zasshi, pp. 641-648, May 2010. doi: 10.1248/yakushi.130.641.
“UniProt”. https://www.uniprot.org/ (accessed Jan. 02, 2022).
“The European Bioinformatics Institute < EMBL-EBI”. https://www.ebi.ac.uk/ (accessed Jan. 02, 2022).
“T-Coffee Multiple Sequence Alignment Program Using Lalign and Dialign | EBI”. https://www.ebi.ac.uk/Tools/msa/tcoffee/ (accessed Jan. 02, 2022).
I. of B. and P. Chemistry, “Geno3D: Automatic modeling of proteins three-dimensional structure”. https://geno3d-prabi.ibcp.fr/cgi-bin/geno3d_automat.pl?page=/GENO3D/geno3d_home.html (accessed Jan. 02, 2022).
“RasMol and OpenRasMol”. http://www.openrasmol.org/ (accessed Jan. 02, 2022).
“PSIPRED Workbench”. http://bioinf.cs.ucl.ac.uk/psipred/ (accessed Jan. 02, 2022).
“Life Sciences and Material Sciences | BIOVIA - Dassault Systèmes”. https://www.3ds.com/products-services/biovia/ (accessed Jan. 02, 2022).
“ProSA-web - Protein structure analysis”. https://prosa.services.came.sbg.ac.at/prosa.php (accessed Jan. 02, 2022).
D. Eisenberg, R. Lüthy, and J. U. Bowie, “[20] Verify3D: Assessment of protein models with three-dimensional profiles”. Methods in Enzymology, vol. 277, pp. 396-404, Jan. 1997, doi: 10.1016/S0076-6879(97)77022-8.
“MOTIF: Searching protein sequence motifs”. https://www.genome.jp/tools/motif/ (accessed Jan. 02, 2022).
“Phylogeny.fr: Home”. https://www.phylogeny.fr/ (accessed Jan. 02, 2022).
“ExPASy - Compute pI_Mw tool”. https://web.expasy.org/compute_pi/
“RCSB PDB: Homepage”. https://www.rcsb.org/ (accessed Jan. 02, 2022).
G. N. Ramachandran, C. Ramakrishnan, and V. Sasisekharan, “Stereochemistry of polypeptide chain configurations”, Journal of Molecular Biology, vol. 7, no. 1. J Mol Biol, pp. 95-99, 1963. doi: 10.1016/S0022-2836(63)80023-6.
“SAVESv6.0 - Structure validation server”. https://saves.mbi.ucla.edu/ (accessed Jan. 02, 2022).
S. Bhaduri, R. L. Buchanan, and J. G. Phillips, “Expanded response surface model for predicting the effects of temperatures, pH, sodium chloride contents and sodium nitrite concentrations on the growth rate of Yersinia enterocolitica”, Journal of Applied Bacteriology, vol. 79, no. 2, pp. 163-170, Aug. 1995, doi: 10.1111/j.1365-2672.1995.tb00930.x.
J. Lafrance-Vanasse, M. Lefebvre, P. di Lello, J. Sygusch, and J. G. Omichinski, “Crystal structures of the organomercurial lyase MerB in its free and mercury-bound forms Insights into the mechanism of methylmercury degradation”, Journal of Biological Chemistry, vol. 284, no. 2, pp. 938-944, 2008, doi: 10.1074/jbc.M807143200.
Y. Nakahara et al., “Purification and characterization of silkworm hemocytes by flow cytometry”, Dev Comp Immunol, vol. 33, no. 4, pp. 439-448, Apr. 2009, doi: 10.1016/J.DCI.2008.09.005.
I. Letunic and P. Bork, “Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation”, Bioinformatics, vol. 23, no. 1, pp. 127-128, Jan. 2007, doi: 10.1093/bioinformatics/btl529.
J. G. Melnick and G. Parkin, “Cleaving mercury-alkyl bonds: A functional model for mercury detoxification by MerB”, Science (1979), vol. 317, no. 5835, pp. 225-227, Jul. 2007, doi: 10.1126/science.1144314.
H. M. Wahba, M. J. Stevenson, A. Mansour, J. Sygusch, D. E. Wilcox, and J. G. Omichinski, “Structural and biochemical characterization of organotin and organolead compounds binding to the organomercurial lyase MerB provide new insights into its mechanism of carbon-metal bond cleavage”, J Am Chem Soc, vol. 139, no. 2, pp. 910-921, Jan. 2017, doi: 10.1021/Jacs.6b11327.
Cómo citar
IEEE
ACM
ACS
APA
ABNT
Chicago
Harvard
MLA
Turabian
Vancouver
Descargar cita
Licencia
Derechos de autor 2022 Kerynd Barona Duque

Esta obra está bajo una licencia internacional Creative Commons Atribución 4.0.
Los autores/as conservarán sus derechos de autor y garantizarán a la revista el derecho de primera publicación de su obra, el cuál estará simultáneamente sujeto a la Licencia de reconocimiento de Creative Commons (CC. Atribución 4.0) que permite a terceros compartir la obra siempre que se indique su autor y su primera publicación en esta revista.
Los autores/as podrán adoptar otros acuerdos de licencia no exclusiva de distribución de la versión de la obra publicada (p. ej.: depositarla en un archivo telemático institucional o publicarla en un volumen monográfico) siempre que se indique la publicación inicial en esta revista.
Se permite y recomienda a los autores/as difundir su obra a través de Internet (p. ej.: en archivos telemáticos institucionales o en su página web) antes y durante el proceso de envío, lo cual puede producir intercambios interesantes y aumentar las citas de la obra publicada. (Véase El efecto del acceso abierto).